Job Results:
Ligand
Structure
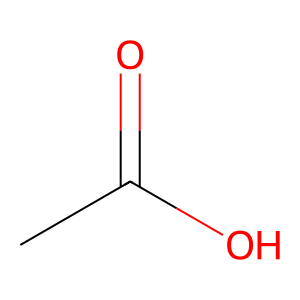
Job ID
60cfc8edcb03d90540341cd2f0373921
Job name
NA
Time
2026-03-02 09:52:38
| Rank | Target | PDB ID |
AirScore |
Detail |
|---|---|---|---|---|
| 1 | Ecto-5'-nucleotidase (CD73) | 4H2G | 4.45 | |
Target general information Gen name NT5E Organism Homo sapiens (Human) Uniprot ID TTD ID Synonyms NT5; CD73 antigen; 5'-nucleotidase; 5'-NT Protein family 5'-nucleotidase family Biochemical class Phosphoric monoester hydrolase Function Exhibits AMP-, NAD-, and NMN-nucleosidase activities. Hydrolyzes extracellular nucleotides into membrane permeable nucleosides. Related diseases Calcification of joints and arteries (CALJA) [MIM:211800]: A condition characterized by adult-onset calcification of the lower extremity arteries, including the iliac, femoral and tibial arteries, and hand and foot capsule joints. Age of onset has been reported as early as the second decade of life, usually involving intense joint pain or calcification in the hands. {ECO:0000269|PubMed:21288095, ECO:0000269|PubMed:24887587}. The disease is caused by variants affecting the gene represented in this entry. Drugs (DrugBank ID) DB00987; DB00806 Interacts with Q9Y225-2; Q8WWF5 EC number EC 3.1.3.5 Uniprot keywords 3D-structure; Alternative splicing; Cell membrane; Direct protein sequencing; Disease variant; Disulfide bond; Glycoprotein; GPI-anchor; Hydrolase; Lipoprotein; Membrane; Metal-binding; Nucleotide-binding; Proteomics identification; Reference proteome; Signal; Zinc Protein physicochemical properties Chain ID A Molecular weight (Da) 24417.6 Length 219 Aromaticity 0.09 Instability index 40.43 Isoelectric point 5.49 Charge (pH=7) -5.75 3D Binding mode Sequence LDDYSTQELGKTIVYLDGSSQSCRFRECNMGNLICDAMINNNLRHADEMFWNHVSMCILNGGGIRSPIDERNDGTITWENLAAVLPFGGTFDLVQLKGSTLKKAFEHSVHRYGQSTGEFLQVGGIHVVYDLSRKPGDRVVKLDVLCTACAVPSYDPLKMDEVYKVILPNFLANGGDGFQMIKDELLRHDSGDQDINVVSTYISKMKVIYPAVEGRIKFS Hydrogen bonds contact Hydrophobic contact | ||||
| 2 | Plasmodium Adenylosuccinate synthetase (Malaria Adss) | 1P9B | 4.45 | |
Target general information Gen name Malaria Adss Organism Plasmodium falciparum Uniprot ID TTD ID Synonyms IMP--aspartate ligase; Adenylosuccinate synthase; AdSS; AMPSase Protein family Adenylosuccinate synthetase family Biochemical class Carbon-nitrogen ligase Function Plays an important role in the salvage pathway for purine nucleotide biosynthesis. Catalyzes the first committed step in the biosynthesis of AMP from IMP. Related diseases Hypertension and brachydactyly syndrome (HTNB) [MIM:112410]: A syndrome characterized by brachydactyly type E, severe salt-independent but age-dependent hypertension, an increased fibroblast growth rate, neurovascular contact at the rostral-ventrolateral medulla, and altered baroreflex blood pressure regulation. It results in death from stroke before age 50 years when untreated. Brachydactyly type E is characterized by shortening of the fingers mainly in the metacarpals and metatarsals. {ECO:0000269|PubMed:25961942}. The disease is caused by variants affecting the gene represented in this entry. Drugs (DrugBank ID) DB03510; DB04315; DB02109 Interacts with NA EC number EC 6.3.4.4 Uniprot keywords 3D-structure; Cytoplasm; GTP-binding; Ligase; Magnesium; Metal-binding; Nucleotide-binding; Purine biosynthesis Protein physicochemical properties Chain ID A Molecular weight (Da) 47877.9 Length 424 Aromaticity 0.09 Instability index 31.72 Isoelectric point 7.63 Charge (pH=7) 1.58 3D Binding mode Sequence GNVVAILGAQWGDEGKGKIIDMLSEYSDITCRFNGGANAGHTISVNDKKYALHLLPCGVLYDNNISVLGNGMVIHVKSLMEEIESVGGKLLDRLYLSNKAHILFDIHQIIDSIQETKKLKEGKQIGTTKRGIGPCYSTKASRIGIRLGTLKNFENFKNMYSKLIDHLMDLYNITEYDKEKELNLFYNYHIKLRDRIVDVISFMNTNLENNKKVLIEGANAAMLDIDFGTYPYVTSSCTTVGGVFSGLGIHHKKLNLVVGVVKSYLTRVGCGPFLTELNNDVGQYLREKGHEYGTTTKRPRRCGWLDIPMLLYVKCINSIDMINLTKLDVLSGLEEILLCVNFKNKKTGELLEKGCYPVEEEISEEYEPVYEKFSGWKEDISTCNEFDELPENAKKYILAIEKYLKTPIVWIGVGPNRKNMIVKK Hydrogen bonds contact Hydrophobic contact | ||||
| 3 | Peptidyl-prolyl cis-trans isomerase G | 2GW2 | 4.44 | |
Target general information Gen name PPIG Organism Homo sapiens (Human) Uniprot ID TTD ID NA Synonyms NA Protein family NA Biochemical class Isomerase Function Cyclosporin A binding.Peptidyl-prolyl cis-trans isomerase activity.RNA binding. Related diseases Intellectual developmental disorder, autosomal dominant 6, with or without seizures (MRD6) [MIM:613970]: A disorder characterized by significantly below average general intellectual functioning associated with impairments in adaptive behavior and manifested during the developmental period. MRD6 additional features may include seizures, hypotonia, abnormal movements, such as dystonia, and autistic features. {ECO:0000269|PubMed:20890276, ECO:0000269|PubMed:23033978, ECO:0000269|PubMed:23160955, ECO:0000269|PubMed:24863970, ECO:0000269|PubMed:25356899, ECO:0000269|PubMed:27839871, ECO:0000269|PubMed:28095420, ECO:0000269|PubMed:38538865}. The disease is caused by variants affecting the gene represented in this entry.; DISEASE: Developmental and epileptic encephalopathy 27 (DEE27) [MIM:616139]: A form of epileptic encephalopathy, a heterogeneous group of severe early-onset epilepsies characterized by refractory seizures, neurodevelopmental impairment, and poor prognosis. Development is normal prior to seizure onset, after which cognitive and motor delays become apparent. {ECO:0000269|PubMed:24272827, ECO:0000269|PubMed:27839871, ECO:0000269|PubMed:27864847, ECO:0000269|PubMed:38538865}. The disease is caused by variants affecting the gene represented in this entry.; DISEASE: A chromosomal aberrations involving GRIN2B has been found in patients with intellectual disability. Translocations t(9;12)(p23;p13.1) and t(10;12)(q21.1;p13.1) with a common breakpoint in 12p13.1. Drugs (DrugBank ID) DB00172 Interacts with Q8N7W2-2; Q8NHQ1; O75553; Q9UI36-2; Q96C98; Q8NC69; P17931; Q6NVH9; Q15365; Q9UL42; Q96CD2; Q14498; Q16637; Q12800; Q9NVV9; PRO_0000037309 [P0C6X7] EC number 5.2.1.8 Uniprot keywords 3D-structure; Alternative splicing; Isomerase; Isopeptide bond; Nucleus; Phosphoprotein; Proteomics identification; Reference proteome; Rotamase; Ubl conjugation Protein physicochemical properties Chain ID A Molecular weight (Da) 19125.4 Length 173 Aromaticity 0.1 Instability index 26.46 Isoelectric point 7.14 Charge (pH=7) 0.24 3D Binding mode Sequence RPRCFFDIAINNQPAGRVVFELFSDVCPKTCENFRCLCTGEKGTGKSTQKPLHYKSCLFHRVVKDFMVQGGDFSEGNGRGGESIYGGFFEDESFAVKHNAAFLLSMANRGKDTNGSQFFITTKPTPHLDGHHVVFGQVISGQEVVREIENQKTDAASKPFAEVRILSCGELIP Hydrogen bonds contact Hydrophobic contact | ||||
| 4 | Phosphoribosylaminoimidazolecarboxamide formyltransferase (ATIC) | 1P4R | 4.44 | |
Target general information Gen name ATIC Organism Homo sapiens (Human) Uniprot ID TTD ID Synonyms PURH; OK/SW-cl.86; Bifunctional purine biosynthesis protein PURH Protein family PurH family Biochemical class Methyltransferase Function Bifunctional enzyme that catalyzes 2 steps in purine biosynthesis. Related diseases AICA-ribosuria due to ATIC deficiency (AICAR) [MIM:608688]: A neurologically devastating inborn error of purine biosynthesis. Patients excrete massive amounts of AICA-riboside in the urine and accumulate AICA-ribotide and its derivatives in erythrocytes and fibroblasts. Clinical features include profound intellectual disability, epilepsy, dysmorphic features and congenital blindness. AICAR inheritance is autosomal recessive. {ECO:0000269|PubMed:15114530}. The disease is caused by variants affecting the gene represented in this entry. Drugs (DrugBank ID) DB02309; DB03442; DB01700; DB01972; DB00563; DB04057; DB00642; DB00116 Interacts with NA EC number NA Uniprot keywords 3D-structure; Acetylation; Alternative splicing; Cytoplasm; Direct protein sequencing; Disease variant; Epilepsy; Hydrolase; Intellectual disability; Multifunctional enzyme; Proteomics identification; Purine biosynthesis; Reference proteome; Transferase Protein physicochemical properties Chain ID A,B Molecular weight (Da) 128556 Length 1177 Aromaticity 0.07 Instability index 38.21 Isoelectric point 6.28 Charge (pH=7) -7.98 3D Binding mode Sequence GQLALFSVSDKTGLVEFARNLTALGLNLVASGGTAKALRDAGLAVRDVSELTGFPEMLGGRVKTLHPAVHAGILARNIPEDNADMARLDFNLIRVVACNLYPFVKTVASPGVTVEEAVEQIDIGGVTLLRAAAKNHARVTVVCEPEDYVVVSTEMQSSESKDTSLETRRQLALKAFTHTAQYDEAISDYFRKQYSKGVSQMPLRYGMNPHQTPAQLYTLQPKLPITVLNGAPGFINLCDALNAWQLVKELKEALGIPAAASFKHVSPAGAAVGIPLSEDEAKVCMVYDLYKTLTPISAAYARARGADRMSSFGDFVALSDVCDVPTAKIISREVSDGIIAPGYEEEALTILSKKKNGNYCVLQMDQSYKPDENEVRTLFGLHLSQKRNNGVVDKSLFSNVVTKNKDLPESALRDLIVATIAVKYTQSNSVCYAKNGQVIGIGAGQQSRIHCTRLAGDKANYWWLRHHPQVLSMKFKTGVKRAEISNAIDQYVTGTIGEDEDLIKWKALFEEVPELLTEAEKKEWVEKLTEVSISSDAFFPFRDNVDRAKRSGVAYIAAPSGSAADKVVIEACDELGIILAHTNLRLFHHQLALFSVSDKTGLVEFARNLTALGLNLVASGGTAKALRDAGLAVRDVSELTGFPEMLGGRVKTLHPAVHAGILARNIPEDNADMARLDFNLIRVVACNLYPFVKTVASPGVTVEEAVEQIDIGGVTLLRAAAKNHARVTVVCEPEDYVVVSTEMQSSESKDTSLETRRQLALKAFTHTAQYDEAISDYFRKQYSKGVSQMPLRYGMNPHQTPAQLYTLQPKLPITVLNGAPGFINLCDALNAWQLVKELKEALGIPAAASFKHVSPAGAAVGIPLSEDEAKVCMVYDLYKTLTPISAAYARARGADRMSSFGDFVALSDVCDVPTAKIISREVSDGIIAPGYEEEALTILSKKKNGNYCVLQMDQSYKPDENEVRTLFGLHLSQKRNNGVVDKSLFSNVVTKNKDLPESALRDLIVATIAVKYTQSNSVCYAKNGQVIGIGAGQQSRIHCTRLAGDKANYWWLRHHPQVLSMKFKTGVKRAEISNAIDQYVTGTIGEDEDLIKWKALFEEVPELLTEAEKKEWVEKLTEVSISSDAFFPFRDNVDRAKRSGVAYIAAPSGSAADKVVIEACDELGIILAHTNLRLFHH Hydrogen bonds contact Hydrophobic contact | ||||
| 5 | Dystroglycan (DAG1) | 7E9K | 4.44 | |
Target general information Gen name DAG1 Organism Homo sapiens (Human) Uniprot ID TTD ID Synonyms Dystrophin-associated glycoprotein 1; Alpha-dystroglycan Protein family NA Biochemical class Dystroglycan protein Function The dystroglycan complex is involved in a number of processes including laminin and basement membrane assembly, sarcolemmal stability, cell survival, peripheral nerve myelination, nodal structure, cell migration, and epithelial polarization. Related diseases Muscular dystrophy-dystroglycanopathy limb-girdle C9 (MDDGC9) [MIM:613818]: An autosomal recessive muscular dystrophy showing onset in early childhood, and associated with intellectual disability without structural brain anomalies. {ECO:0000269|PubMed:21388311, ECO:0000269|PubMed:25503980}. The disease is caused by variants affecting the gene represented in this entry. MDDGC7 is caused by DAG1 mutations that interfere with normal post-translational processing, resulting in defective DAG1 glycosylation and impaired interactions with extracellular-matrix components. Other muscular dystrophy-dystroglycanopathies are caused by defects in enzymes involved in protein O-glycosylation.; DISEASE: Muscular dystrophy-dystroglycanopathy congenital with brain and eye anomalies A9 (MDDGA9) [MIM:616538]: An autosomal recessive disorder characterized by congenital muscular dystrophy associated with cobblestone lissencephaly and other brain anomalies, eye malformations, profound intellectual disability, and death usually in the first years of life. Included diseases are the more severe Walker-Warburg syndrome and the slightly less severe muscle-eye-brain disease. {ECO:0000269|PubMed:24052401, ECO:0000269|PubMed:25934851}. The disease is caused by variants affecting the gene represented in this entry. Drugs (DrugBank ID) NA Interacts with P16333; Q3T1J5; PRO_0000021066 [Q14118]; P46939 EC number NA Uniprot keywords 3D-structure; Autocatalytic cleavage; Cell membrane; Congenital muscular dystrophy; Cytoplasm; Cytoskeleton; Disease variant; Disulfide bond; Dystroglycanopathy; Glycoprotein; Host cell receptor for virus entry; Host-virus interaction; Limb-girdle muscular dystrophy; Lissencephaly; Membrane; Nucleus; Phosphoprotein; Postsynaptic cell membrane; Proteomics identification; Receptor; Reference proteome; Secreted; Signal; Synapse; Transmembrane; Transmembrane helix Protein physicochemical properties Chain ID A,C Molecular weight (Da) 62559.7 Length 546 Aromaticity 0.11 Instability index 38.37 Isoelectric point 9 Charge (pH=7) 10.42 3D Binding mode Sequence PAPALRIDYPKALQILTEGGTHMVCTGRTHTDRLCRFKWLCYSSEAEEFIFFHGNASVMLPSLGSRRFQPALLDLSTVEDHNTQYFNFVELPAAALRFMPKPVFVPDVALIANRFNPDNLMHVFHDDLLPLFYTLRQFPGLAREARLFFMEGWGEGAHFDLYKLLSPKQPLLRAQLKALGRLLCFSHAFVGLSKVTTWYQYGFVQPQGPKANILVSGNEIRQFAHFLMEKLNVSEEYILVFSRTQNRLILNEAELLLALAQEFQMKTVTVSLEDHAFADVVRLVSNASMLVSMHGAQLVTALFLPRGAAVVELFPYAVNPDHYTPYKTLATLPGMDLQYIAWQNTMPENTVTHPERPWDQGGIAHLDRAEQARILQSREVPRHLCCRNPEWLFRIYQDTKVDIPSLIQTIRRVVKGHPGPRKQKWTVSLYPGKVREARCQASVASEARLSVSWQIPWNLKYLKVREVKYEVWLQEQGENTYVPYMLALQNHTFTENIKPFTTYLVWIRCIFNKTLLGPFADVLVCSTRTRGAIIQTPTLGPIQPTR Hydrogen bonds contact Hydrophobic contact | ||||
| 6 | Plasmodium Adenylosuccinate synthetase (Malaria Adss) | 1P9B | 4.44 | |
Target general information Gen name Malaria Adss Organism Plasmodium falciparum Uniprot ID TTD ID Synonyms IMP--aspartate ligase; Adenylosuccinate synthase; AdSS; AMPSase Protein family Adenylosuccinate synthetase family Biochemical class Carbon-nitrogen ligase Function Plays an important role in the salvage pathway for purine nucleotide biosynthesis. Catalyzes the first committed step in the biosynthesis of AMP from IMP. Related diseases Hypertension and brachydactyly syndrome (HTNB) [MIM:112410]: A syndrome characterized by brachydactyly type E, severe salt-independent but age-dependent hypertension, an increased fibroblast growth rate, neurovascular contact at the rostral-ventrolateral medulla, and altered baroreflex blood pressure regulation. It results in death from stroke before age 50 years when untreated. Brachydactyly type E is characterized by shortening of the fingers mainly in the metacarpals and metatarsals. {ECO:0000269|PubMed:25961942}. The disease is caused by variants affecting the gene represented in this entry. Drugs (DrugBank ID) DB03510; DB04315; DB02109 Interacts with NA EC number EC 6.3.4.4 Uniprot keywords 3D-structure; Cytoplasm; GTP-binding; Ligase; Magnesium; Metal-binding; Nucleotide-binding; Purine biosynthesis Protein physicochemical properties Chain ID A Molecular weight (Da) 47877.9 Length 424 Aromaticity 0.09 Instability index 31.72 Isoelectric point 7.63 Charge (pH=7) 1.58 3D Binding mode Sequence GNVVAILGAQWGDEGKGKIIDMLSEYSDITCRFNGGANAGHTISVNDKKYALHLLPCGVLYDNNISVLGNGMVIHVKSLMEEIESVGGKLLDRLYLSNKAHILFDIHQIIDSIQETKKLKEGKQIGTTKRGIGPCYSTKASRIGIRLGTLKNFENFKNMYSKLIDHLMDLYNITEYDKEKELNLFYNYHIKLRDRIVDVISFMNTNLENNKKVLIEGANAAMLDIDFGTYPYVTSSCTTVGGVFSGLGIHHKKLNLVVGVVKSYLTRVGCGPFLTELNNDVGQYLREKGHEYGTTTKRPRRCGWLDIPMLLYVKCINSIDMINLTKLDVLSGLEEILLCVNFKNKKTGELLEKGCYPVEEEISEEYEPVYEKFSGWKEDISTCNEFDELPENAKKYILAIEKYLKTPIVWIGVGPNRKNMIVKK Hydrogen bonds contact Hydrophobic contact | ||||
| 7 | Ubiquitin carboxyl-terminal hydrolase 2 (USP2) | 5XU8 | 4.44 | |
Target general information Gen name USP2 Organism Homo sapiens (Human) Uniprot ID TTD ID Synonyms Ubiquitin-specific-processing protease 2; Ubiquitin thioesterase 2; UBP41; Deubiquitinating enzyme 2; 41 kDa ubiquitin-specific protease Protein family Peptidase C19 family, USP2 subfamily Biochemical class Peptidase Function Isoform 1 and isoform 4 possess both ubiquitin-specific peptidase and isopeptidase activities. Deubiquitinates MDM2 without reversing MDM2-mediated p53/TP53 ubiquitination and thus indirectly promotes p53/TP53 degradation and limits p53 activity. Has no deubiquitinase activity against p53/TP53. Prevents MDM2-mediated degradation of MDM4. Plays a role in the G1/S cell-cycle progression in normal and cancer cells. Regulates the circadian clock by modulating its intrinsic circadian rhythm and its capacity to respond to external cues. Associates with clock proteins and deubiquitinates core clock component PER1 but does not affect its overall stability. Regulates the nucleocytoplasmic shuttling and nuclear retention of PER1 and its repressive role on the clock transcription factors CLOCK and ARNTL/BMAL1. Plays a role in the regulation of myogenic differentiation of embryonic muscle cells. Hydrolase that deubiquitinates polyubiquitinated target proteins such as MDM2, MDM4 and CCND1. Related diseases Defects in AKT2 are a cause of susceptibility to breast cancer (BC). AKT2 promotes metastasis of tumor cells without affecting the latency of tumor development. May play a role in glioblastoma cell survival (PubMed:20167810). {ECO:0000269|PubMed:20167810}.; DISEASE: Type 2 diabetes mellitus (T2D) [MIM:125853]: A multifactorial disorder of glucose homeostasis caused by a lack of sensitivity to insulin. Affected individuals usually have an obese body habitus and manifestations of a metabolic syndrome characterized by diabetes, insulin resistance, hypertension and hypertriglyceridemia. The disease results in long-term complications that affect the eyes, kidneys, nerves, and blood vessels. {ECO:0000269|PubMed:15166380, ECO:0000269|PubMed:19164855}. Disease susceptibility is associated with variants affecting the gene represented in this entry.; DISEASE: Hypoinsulinemic hypoglycemia with hemihypertrophy (HIHGHH) [MIM:240900]: A disorder characterized by hypoglycemia, low insulin levels, low serum levels of ketone bodies and branched-chain amino acids, left-sided hemihypertrophy, neonatal macrosomia, reduced consciousness and hypoglycemic seizures. {ECO:0000269|PubMed:21979934}. The disease is caused by variants affecting the gene represented in this entry. Drugs (DrugBank ID) NA Interacts with Q9NYB9-2; P12814; P35609; Q08043; Q86U10; Q86V38; P56945; Q8TD16-2; Q96CA5; A2RRN7; Q13137; Q9H257-2; Q96JN2-2; Q2TAC2; A6NC98; Q96MT8-3; Q8NHQ1; Q9BSW2; Q8N4Y2-3; Q8WTU0; O75140-2; Q9NRI5-2; Q8N9I9; Q9H596; Q8WWB3; Q5JST6; Q9NRA8; O00471; Q96B26; P57678; Q08379; Q9NYA3; A6NEM1; Q6PI77; Q14451-3; Q4V328; Q9NSC5; Q9UJC3; Q96ED9-2; Q8IYA8; Q9UKT9; Q5TA45; Q96N16; O75564-2; Q674X7-2; Q9BVG8; Q9BVG8-5; P19012; Q7Z3Y8; Q15323; Q14525; O76011; Q92764; Q6A162; Q9UBR4-2; Q969G2; Q03252; Q9BRK4; Q00987; Q9UJV3-2; Q5VZ52; Q13084; Q5JR59; Q5JR59-3; Q15742; Q9GZM8; I6L9F6; P07196; O43482; Q96CV9; Q4G0R1; Q9NRD5; Q58EX7; Q8ND90; Q16633; Q9GZV8; Q6MZQ0; Q15276; Q8HWS3; Q59EK9-3; P60903; O14492-2; O60504; Q99932-2; A6NLX3; P51692; Q86VP1; Q8WW24; Q9UBB9; Q08117-2; Q03169; Q13077; Q12933; Q9Y4K3; P36406; P14373; Q86XT4; Q15654; Q8N6Y0; Q70EL1-9; Q9UK41-2; Q8N1B4; O96006; Q9NZV7; Q9UGI0; P05067; P54253; G5E9A7; Q01658; Q00403; Q9Y5Q9; P04792; O43464; P42858; Q8WXH2; O60333-2; A0A6Q8PF08; O60260-5; P60891; Q9Y3C5; Q7Z333; P37840; P00441; Q7Z699; Q13148; O76024 EC number EC 3.4.19.12 Uniprot keywords 3D-structure; Alternative splicing; Biological rhythms; Cell cycle; Cytoplasm; Hydrolase; Membrane; Metal-binding; Myogenesis; Nucleus; Protease; Proteomics identification; Reference proteome; Thiol protease; Ubl conjugation pathway; Zinc Protein physicochemical properties Chain ID A Molecular weight (Da) 37785.5 Length 327 Aromaticity 0.11 Instability index 42.45 Isoelectric point 8.23 Charge (pH=7) 3.56 3D Binding mode Sequence QGLAGLRNLGNTCFMNSILQCLSNTRELRDYCLQRLYMRDLHHGSNAHTALVEEFAKLIQTIWTSSPNDVVSPSEFKTQIQRYAPRFVGYNQQDAQEFLRFLLDGLHNEVNRVNLDHLPDDEKGRQMWRKYLEREDSRIGDLFVGQLKSSLTCTDCGYCSTVFDPFWDLSLPIAKRGYPEVTLMDCMRLFTKEDVLDGDEKPTCCRCRGRKRCIKKFSIQRFPKILVLHLKRFSESRIRTSKLTTFVNFPLRDLDLREFASENTNHAVYNLYAVSNHSGTTMGGHYTAYCRSPGTGEWHTFNDSSVTPMSSSQVRTSDAYLLFYELA Hydrogen bonds contact Hydrophobic contact | ||||
| 8 | Vitamin D3 receptor (VDR) | 3B0T | 4.43 | |
Target general information Gen name VDR Organism Homo sapiens (Human) Uniprot ID TTD ID Synonyms Vitamin D(3) receptor; Nuclear vitamin D receptor; Nuclear receptor subfamily 1 group I member 1; NR1I1; 1,25-dihydroxyvitamin D3 receptor Protein family Nuclear hormone receptor family, NR1 subfamily Biochemical class Nuclear hormone receptor Function Enters the nucleus upon vitamin D3 binding where it forms heterodimers with the retinoid X receptor/RXR. The VDR-RXR heterodimers bind to specific response elements on DNA and activate the transcription of vitamin D3-responsive target genes. Plays a central role in calcium homeostasis. Nuclear receptor for calcitriol, the active form of vitamin D3 which mediates the action of this vitamin on cells. Related diseases Rickets vitamin D-dependent 2A (VDDR2A) [MIM:277440]: A disorder of vitamin D metabolism resulting in severe rickets, hypocalcemia and secondary hyperparathyroidism. Most patients have total alopecia in addition to rickets. {ECO:0000269|PubMed:1652893, ECO:0000269|PubMed:17970811, ECO:0000269|PubMed:2177843, ECO:0000269|PubMed:2849209, ECO:0000269|PubMed:28698609, ECO:0000269|PubMed:7828346, ECO:0000269|PubMed:8106618, ECO:0000269|PubMed:8381803, ECO:0000269|PubMed:8392085, ECO:0000269|PubMed:8675579, ECO:0000269|PubMed:8961271, ECO:0000269|PubMed:9005998}. The disease is caused by variants affecting the gene represented in this entry. Drugs (DrugBank ID) DB07530; DB08742; DB01436; DB04891; DB00146; DB02300; DB00136; DB00169; DB04540; DB05024; DB11672; DB14635; DB01070; DB06410; DB05295; DB06194; DB00153; DB04796; DB03451; DB00910; DB04258; DB11094 Interacts with P35222; Q09472; Q15648; P50222; Q15788; P26045; P19793; Q13573; Q13501; P04637; Q15645; Q9JLI4; P28700; X5D778; Q96HA8; Q01804; Q96S38; P48443 EC number NA Uniprot keywords 3D-structure; Alternative splicing; Cytoplasm; Disease variant; DNA-binding; Metal-binding; Nucleus; Proteomics identification; Receptor; Reference proteome; Transcription; Transcription regulation; Ubl conjugation; Zinc; Zinc-finger Protein physicochemical properties Chain ID A Molecular weight (Da) 28781 Length 254 Aromaticity 0.07 Instability index 47.69 Isoelectric point 6.15 Charge (pH=7) -3.44 3D Binding mode Sequence ALRPKLSEEQQRIIAILLDAHHKTYDPTYSDFCQFRPPVRVNDGGGSVTLELSQLSMLPHLADLVSYSIQKVIGFAKMIPGFRDLTSEDQIVLLKSSAIEVIMLRSNESFTMDDMSWTCGNQDYKYRVSDVTKAGHSLELIEPLIKFQVGLKKLNLHEEEHVLLMAICIVSPDRPGVQDAALIEAIQDRLSNTLQTYIRCRHPPPGSHLLYAKMIQKLADLRSLNEEHSKQYRCLSFQPECSMKLTPLVLEVFG Hydrogen bonds contact Hydrophobic contact | ||||
| 9 | 3-ketoacyl-CoA thiolase, peroxisomal | 2IIK | 4.43 | |
Target general information Gen name ACAA1 Organism Homo sapiens (Human) Uniprot ID TTD ID NA Synonyms PTHIO;ACAA Protein family Thiolase-like superfamily, Thiolase family Biochemical class Transferase Function Acetate CoA-transferase activity.Acetyl-CoA C-acyltransferase activity.Palmitoyl-CoA oxidase activity. Related diseases Factor VII deficiency (FA7D) [MIM:227500]: A hemorrhagic disease with variable presentation. The clinical picture can be very severe, with the early occurrence of intracerebral hemorrhages or repeated hemarthroses, or, in contrast, moderate with cutaneous-mucosal hemorrhages (epistaxis, menorrhagia) or hemorrhages provoked by a surgical intervention. Finally, numerous subjects are completely asymptomatic despite very low factor VII levels. {ECO:0000269|PubMed:10862079, ECO:0000269|PubMed:11091194, ECO:0000269|PubMed:11129332, ECO:0000269|PubMed:12472587, ECO:0000269|PubMed:14717781, ECO:0000269|PubMed:1634227, ECO:0000269|PubMed:18976247, ECO:0000269|PubMed:19432927, ECO:0000269|PubMed:19751712, ECO:0000269|PubMed:2070047, ECO:0000269|PubMed:21206266, ECO:0000269|PubMed:21372693, ECO:0000269|PubMed:26761581, ECO:0000269|PubMed:7974346, ECO:0000269|PubMed:7981691, ECO:0000269|PubMed:8043443, ECO:0000269|PubMed:8204879, ECO:0000269|PubMed:8242057, ECO:0000269|PubMed:8364544, ECO:0000269|PubMed:8652821, ECO:0000269|PubMed:8844208, ECO:0000269|PubMed:8883260, ECO:0000269|PubMed:8940045, ECO:0000269|PubMed:9414278, ECO:0000269|PubMed:9452082, ECO:0000269|PubMed:9576180}. The disease is caused by variants affecting the gene represented in this entry. Drugs (DrugBank ID) NA Interacts with Q9BSK4; Q12800 EC number 2.3.1.155; 2.3.1.16; 2.3.1.9 Uniprot keywords 3D-structure; Acyltransferase; Alternative splicing; Fatty acid metabolism; Lipid metabolism; Peroxisome; Phosphoprotein; Proteomics identification; Reference proteome; Transferase; Transit peptide Protein physicochemical properties Chain ID A,B Molecular weight (Da) 37940.2 Length 364 Aromaticity 0.04 Instability index 41.27 Isoelectric point 8.59 Charge (pH=7) 3.99 3D Binding mode Sequence DVVVVHGRRTAICRAGRGGFKDTTPDELLSAVMTAVLKDVNLRPEQLGDICVGNVLQPGAGAIMARIAQFLSDIPETVPLSTVNRQCSSGLQAVASIAGGIRNGSYDIGMACGVESMSLAMGITSENVAERFGISREKQDTFALASQQKAARAQSKGCFQAEIVPVTTTVHDDKGTKRSITVTQDEGIRPSTTMEGLAKLKPAFKKDGSTTAGNSSQVSDGAAAILLARRSKAEELGLPILGVLRSYAVVGVPPDIMGIGPAYAIPVALQKAGLTVSDVDIFEINEAFASQAAYCVEKLRLPPEKVNPLGGAVALGHPLGCTGARQVITLLNELKRRGKRAYGVVSMCIGTGMGAAAVFEYPGN Hydrogen bonds contact Hydrophobic contact | ||||
| 10 | Protein-tyrosine phosphatase 1B (PTP1B) | 2F71 | 4.43 | |
Target general information Gen name PTPN1 Organism Homo sapiens (Human) Uniprot ID TTD ID Synonyms Tyrosine-protein phosphatase non-receptor type 1; PTP-1B Protein family Protein-tyrosine phosphatase family, Non-receptor class 1 subfamily Biochemical class Phosphoric monoester hydrolase Function Mediates dephosphorylation of EIF2AK3/PERK; inactivating the protein kinase activity of EIF2AK3/PERK. May play an important role in CKII- and p60c-src-induced signal transduction cascades. May regulate the EFNA5-EPHA3 signaling pathway which modulates cell reorganization and cell-cell repulsion. May also regulate the hepatocyte growth factor receptor signaling pathway through dephosphorylation of MET. Tyrosine-protein phosphatase which acts as a regulator of endoplasmic reticulum unfolded protein response. Related diseases Can contribute to cancer cell survival, proliferation, migration, and invasion, and tumor angiogenesis and metastasis. May contribute to cancer pathogenesis by promoting inflammatory responses and recruitment of tumor-infiltrating macrophages.; DISEASE: Abnormally high expression of soluble isoforms (isoform 2, isoform 3 or isoform 4) may be a cause of preeclampsia. Drugs (DrugBank ID) DB08549; DB08783; DB03483; DB08593; DB04800; DB03670; DB03102; DB02072; DB02622; DB07295; DB04088; DB01820; DB02259; DB03311; DB04142; DB02620; DB07298; DB01734; DB08147; DB03557; DB07197; DB06829; DB07130; DB03714; DB07480; DB02014; DB07730; DB08001; DB07134; DB08591; DB07289; DB08397; DB04001; DB02827; DB07719; DB06887; DB04204; DB02420; DB07263; DB02615; DB03982; DB06521; DB05506; DB08003; DB03661; DB04525; DB02784; DB07651; DB02662; DB08371; DB02977; DB06333; DB02436; DB04285; DB02651; DB03154 Interacts with Q13520; P56945; P11274-1; P07384; Q03135; Q14247; P00533; Q9GZR5; P19235; P10912; P62993; P08069; P06213; P06213-1; P05556; P05106; O60674; O43561; P08581; P04629; Q16288; P09619; P57054; P08922; P12931; P40763; P42229; Q9NPL8; Q96HV5; Q8N661; Q9H1D0; P10599; Q61140; Q63767; P15116; Q63768; P62994; P35570; P05622; P10686; P34152; Q8VI36; Q9WUD9; P63166 EC number EC 3.1.3.48 Uniprot keywords 3D-structure; Acetylation; Direct protein sequencing; Endoplasmic reticulum; Hydrolase; Membrane; Oxidation; Phosphoprotein; Protein phosphatase; Proteomics identification; Reference proteome; S-nitrosylation Protein physicochemical properties Chain ID A Molecular weight (Da) 34541 Length 297 Aromaticity 0.1 Instability index 35.91 Isoelectric point 5.91 Charge (pH=7) -5.37 3D Binding mode Sequence EMEKEFEQIDKSGSWAAIYQDIRHEASDFPCRVAKLPKNKNRNRYRDVSPFDHSRIKLHQEDNDYINASLIKMEEAQRSYILTQGPLPNTCGHFWEMVWEQKSRGVVMLNRVMEKGSLKCAQYWPQKEEKEMIFEDTNLKLTLISEDIKSYYTVRQLELENLTTQETREILHFHYTTWPDFGVPESPASFLNFLFKVRESGSLSPEHGPVVVHCSAGIGRSGTFCLADTCLLLMDKRKDPSSVDIKKVLLEMRKFRMGLIQTADQLRFSYLAVIEGAKFIMGDSSVQDQWKELSHED Hydrogen bonds contact Hydrophobic contact | ||||
| 11 | Peptidyl-prolyl cis-trans isomerase G | 2GW2 | 4.43 | |
Target general information Gen name PPIG Organism Homo sapiens (Human) Uniprot ID TTD ID NA Synonyms NA Protein family NA Biochemical class Isomerase Function Cyclosporin A binding.Peptidyl-prolyl cis-trans isomerase activity.RNA binding. Related diseases Intellectual developmental disorder, autosomal dominant 6, with or without seizures (MRD6) [MIM:613970]: A disorder characterized by significantly below average general intellectual functioning associated with impairments in adaptive behavior and manifested during the developmental period. MRD6 additional features may include seizures, hypotonia, abnormal movements, such as dystonia, and autistic features. {ECO:0000269|PubMed:20890276, ECO:0000269|PubMed:23033978, ECO:0000269|PubMed:23160955, ECO:0000269|PubMed:24863970, ECO:0000269|PubMed:25356899, ECO:0000269|PubMed:27839871, ECO:0000269|PubMed:28095420, ECO:0000269|PubMed:38538865}. The disease is caused by variants affecting the gene represented in this entry.; DISEASE: Developmental and epileptic encephalopathy 27 (DEE27) [MIM:616139]: A form of epileptic encephalopathy, a heterogeneous group of severe early-onset epilepsies characterized by refractory seizures, neurodevelopmental impairment, and poor prognosis. Development is normal prior to seizure onset, after which cognitive and motor delays become apparent. {ECO:0000269|PubMed:24272827, ECO:0000269|PubMed:27839871, ECO:0000269|PubMed:27864847, ECO:0000269|PubMed:38538865}. The disease is caused by variants affecting the gene represented in this entry.; DISEASE: A chromosomal aberrations involving GRIN2B has been found in patients with intellectual disability. Translocations t(9;12)(p23;p13.1) and t(10;12)(q21.1;p13.1) with a common breakpoint in 12p13.1. Drugs (DrugBank ID) DB00172 Interacts with Q8N7W2-2; Q8NHQ1; O75553; Q9UI36-2; Q96C98; Q8NC69; P17931; Q6NVH9; Q15365; Q9UL42; Q96CD2; Q14498; Q16637; Q12800; Q9NVV9; PRO_0000037309 [P0C6X7] EC number 5.2.1.8 Uniprot keywords 3D-structure; Alternative splicing; Isomerase; Isopeptide bond; Nucleus; Phosphoprotein; Proteomics identification; Reference proteome; Rotamase; Ubl conjugation Protein physicochemical properties Chain ID A Molecular weight (Da) 19125.4 Length 173 Aromaticity 0.1 Instability index 26.46 Isoelectric point 7.14 Charge (pH=7) 0.24 3D Binding mode Sequence RPRCFFDIAINNQPAGRVVFELFSDVCPKTCENFRCLCTGEKGTGKSTQKPLHYKSCLFHRVVKDFMVQGGDFSEGNGRGGESIYGGFFEDESFAVKHNAAFLLSMANRGKDTNGSQFFITTKPTPHLDGHHVVFGQVISGQEVVREIENQKTDAASKPFAEVRILSCGELIP Hydrogen bonds contact Hydrophobic contact | ||||
| 12 | Cytochrome P450 3A4 | 4D6Z | 4.42 | |
Target general information Gen name CYP3A4 Organism Homo sapiens (Human) Uniprot ID TTD ID NA Synonyms CYP3A3 Protein family Cytochrome P450 family Biochemical class Oxidoreductase Function 1,8-cineole 2-exo-monooxygenase activity.Albendazole monooxygenase activity.Aromatase activity.Caffeine oxidase activity.Enzyme binding.Estrogen 16-alpha-hydroxylase activity.Heme binding.Iron ion binding.Monooxygenase activity.Oxidoreductase activity.Oxygen binding.Quinine 3-monooxygenase activity.Steroid binding.Steroid hydroxylase activity.Taurochenodeoxycholate 6alpha-hydroxylase activity.Testosterone 6-beta-hydroxylase activity.Vitamin D 24-hydroxylase activity.Vitamin D3 25-hydroxylase activity. Related diseases Vitamin D-dependent rickets 3 (VDDR3) [MIM:619073]: An autosomal dominant disorder of vitamin D metabolism resulting in early-onset rickets, reduced serum levels of the vitamin D metabolites 25-hydroxyvitamin D and 1,25-dihydroxyvitamin D, and deficient responsiveness to parent and activated forms of vitamin D. {ECO:0000269|PubMed:29461981}. The gene represented in this entry is involved in disease pathogenesis. Drugs (DrugBank ID) DB08496; DB14055; DB12537; DB12629; DB01456; DB04070; DB11919; DB12515; DB11932; DB12001; DB05812; DB14973; DB11703; DB01418; DB00316; DB00819; DB15568; DB00546; DB08838; DB00518; DB00240; DB00041; DB04630; DB00802; DB00346; DB09026; DB00918; DB06203; DB00969; DB12015; DB14003; DB00404; DB06403; DB06742; DB13141; DB00288; DB00357; DB01424; DB01223; DB01118; DB00321; DB00381; DB00701; DB01217; DB01536; DB01435; DB11901; DB06605; DB00714; DB05676; DB00673; DB01352; DB09229; DB00278; DB01238; DB14185; DB06413; DB01169; DB06697; DB12597; DB06216; DB00637; DB11586; DB01072; DB16098; DB01076; DB01117; DB15011; DB06237; DB15233; DB06442; DB11995; DB06318; DB06626; DB00972; DB09230; DB04957; DB00207; DB12781; DB13997; DB04975; DB01483; DB11817; DB09227; DB00394; DB08903; DB05015; DB16703; DB15463; DB13488; DB09231; DB00865; DB01244; DB15982; DB00443; DB14669; DB12236; DB00307; DB01393; DB01128; DB11799; DB04794; DB00905; DB13746; DB16536; DB00612; DB13975; DB09223; DB08873; DB00188; DB00559; DB06616; DB07348; DB08870; DB09128; DB12267; DB01194; DB05541; DB01200; DB09017; DB11752; DB01222; DB00297; DB00921; DB00490; DB01008; DB09173; DB06772; DB00248; DB08875; DB00201; DB04886; DB00136; DB08907; DB01152; DB09061; DB14737; DB12218; DB11791; DB08502; DB06774; DB00564; DB11383; DB11960; DB06016; DB13835; DB01136; DB14984; DB06634; DB00520; DB01333; DB00482; DB06119; DB09063; DB00439; DB06419; DB00185; DB06777; DB00446; DB00475; DB13528; DB00608; DB00856; DB01114; DB00477; DB00356; DB00169; DB01410; DB09201; DB09232; DB01166; DB00501; DB01012; DB00568; DB00537; DB00604; DB00215; DB01211; DB12499; DB04920; DB01190; DB00349; DB11750; DB01013; DB13158; DB14652; DB00845; DB00636; DB06470; DB01242; DB01068; DB00575; DB00758; DB13843; DB00628; DB01559; DB00257; DB00363; DB09065; DB05239; DB00907; DB00318; DB01394; DB06342; DB00872; DB00286; DB12483; DB04652; DB01285; DB14681; DB01380; DB13003; DB08865; DB11672; DB14635; DB04838; DB00924; DB00531; DB00091; DB04839; DB00987; DB08912; DB09102; DB11963; DB01764; DB01406; DB11779; DB06292; DB04884; DB11682; DB00250; DB15031; DB00496; DB09234; DB12941; DB01264; DB09183; DB01254; DB00694; DB01609; DB11921; DB11943; DB11637; DB00705; DB13857; DB01151; DB00304; DB01260; DB06780; DB01134; DB06700; DB12161; DB01234; DB14649; DB11487; DB09555; DB05351; DB04856; DB14068; DB00514; DB00647; DB14063; DB11994; DB00829; DB00586; DB00485; DB09123; DB00255; DB09095; DB06781; DB01396; DB11274; DB01551; DB11273; DB13345; DB13385; DB00320; DB00343; DB01093; DB08995; DB13347; DB00954; DB00280; DB00822; DB02520; DB01248; DB00204; DB00757; DB08930; DB01184; DB00843; DB11400; DB12301; DB06446; DB05928; DB00590; DB01142; DB00997; DB00254; DB00470; DB04855; DB01395; DB00476; DB11952; DB00378; DB11742; DB14240; DB01127; DB14598; DB14600; DB00625; DB09235; DB06374; DB11979; DB11574; DB00216; DB15444; DB09039; DB09101; DB14064; DB13874; DB11718; DB13007; DB11986; DB08899; DB08992; DB00751; DB00668; DB00700; DB12266; DB01873; DB11405; DB03515; DB02187; DB12329; DB12147; DB01049; DB01253; DB00696; DB00530; DB00199; DB01175; DB11823; DB14575; DB09119; DB00736; DB01215; DB09381; DB12235; DB00783; DB13952; DB13953; DB13954; DB13955; DB13956; DB01196; DB00655; DB04574; DB00402; DB00330; DB00898; DB00977; DB00593; DB08794; DB01466; DB00823; DB09166; DB00294; DB00773; DB01628; DB14766; DB06414; DB13866; DB01590; DB00990; DB00973; DB12500; DB00949; DB01023; DB08980; DB00574; DB00813; DB06702; DB12265; DB08874; DB01216; DB16165; DB13961; DB04908; DB00301; DB00196; DB00687; DB00663; DB04841; DB00180; DB01544; DB00591; DB01047; DB08971; DB00324; DB00472; DB08970; DB14634; DB09378; DB14637; DB00846; DB00690; DB13338; DB04842; DB00499; DB13867; DB08906; DB00588; DB01095; DB00176; DB12307; DB08905; DB01319; DB06717; DB14019; DB01320; DB12010; DB11796; DB11679; DB00947; DB02703; DB15149; DB00674; DB12923; DB05087; DB00317; DB01241; DB01645; DB12184; DB06730; DB11619; DB12141; DB01381; DB11978; DB13879; DB00143; DB01016; DB08909; DB00986; DB05814; DB00889; DB10534; DB11575; DB00365; DB00400; DB01018; DB06786; DB01218; DB13728; DB00502; DB01159; DB05212; DB01275; DB00956; DB00769; DB00741; DB14538; DB14539; DB14540; DB14541; DB14542; DB14543; DB14545; DB14544; DB01611; DB14570; DB06789; DB00557; DB12471; DB09053; DB01050; DB11737; DB09054; DB01181; DB04946; DB00619; DB09262; DB00458; DB00724; DB05039; DB08953; DB00808; DB00224; DB06370; DB11886; DB13293; DB01029; DB00762; DB11633; DB06636; DB00951; DB00982; DB00270; DB11757; DB01167; DB09083; DB08820; DB00602; DB14568; DB04845; DB09570; DB01221; DB01587; DB06738; DB01026; DB09309; DB05903; DB09236; DB06218; DB06791; DB00448; DB01259; DB06685; DB14723; DB12825; DB11951; DB15673; DB16217; DB09078; DB00528; DB11560; DB06469; DB12070; DB01006; DB01227; DB09237; DB01002; DB06282; DB05667; DB00825; DB08918; DB00367; DB00281; DB13766; DB08882; DB17083; DB01583; DB00589; DB09198; DB14065; DB08827; DB01206; DB06448; DB16222; DB00836; DB01601; DB00455; DB00186; DB04871; DB12130; DB09195; DB12089; DB00678; DB14596; DB00227; DB09212; DB08933; DB09280; DB06077; DB06708; DB08815; DB12674; DB12474; DB04829; DB13074; DB08932; DB09238; DB16226; DB04835; DB06234; DB14921; DB00643; DB14009; DB09124; DB00603; DB00253; DB00358; DB00351; DB11529; DB14659; DB00814; DB00170; DB00454; DB09383; DB01071; DB01357; DB04817; DB00333; DB04833; DB00763; DB00563; DB01028; DB09241; DB00353; DB00959; DB14644; DB12952; DB06710; DB00247; DB01233; DB00264; DB00916; DB01011; DB15489; DB00379; DB06148; DB01388; DB01110; DB00683; DB13456; DB06595; DB00834; DB04896; DB13287; DB08893; DB11792; DB00370; DB12489; DB16236; DB06587; DB00648; DB01204; DB16390; DB00745; DB11763; DB00764; DB14512; DB00471; DB00295; DB09205; DB00688; DB01024; DB11605; DB00486; DB14011; DB00607; DB12092; DB11691; DB06230; DB09049; DB01183; DB00731; DB04861; DB01149; DB00220; DB11828; DB09199; DB09048; DB00238; DB00627; DB00622; DB02701; DB00184; DB01115; DB09239; DB04868; DB09240; DB06712; DB04743; DB00393; DB09079; DB16691; DB12005; DB00401; DB01595; DB01054; DB00435; DB11636; DB13981; DB06713; DB14678; DB00717; DB09371; DB01059; DB00957; DB09389; DB00540; DB06174; DB06152; DB00104; DB06670; DB00334; DB09074; DB11442; DB14881; DB00768; DB16267; DB12513; DB09568; DB00338; DB00904; DB11130; DB04911; DB01083; DB01173; DB11837; DB09330; DB04938; DB13500; DB00776; DB12532; DB00239; DB01062; DB00497; DB06412; DB01192; DB12612; DB01229; DB11697; DB09073; DB01267; DB00377; DB05467; DB06603; DB00213; DB00617; DB01384; DB08439; DB00910; DB09297; DB00715; DB06663; DB03010; DB06589; DB00082; DB15102; DB13791; DB00312; DB11198; DB08883; DB01186; DB01074; DB08922; DB00850; DB12978; DB03783; DB00780; DB01174; DB00946; DB00191; DB00812; DB00252; DB13878; DB01085; DB05316; DB00337; DB01100; DB06762; DB09090; DB01132; DB13941; DB12582; DB01621; DB04951; DB17472; DB11642; DB04977; DB12240; DB08910; DB08901; DB12016; DB01263; DB05478; DB15822; DB01411; DB06209; DB01588; DB01058; DB01130; DB00860; DB15566; DB14633; DB14631; DB00635; DB14646; DB13208; DB02789; DB04825; DB05154; DB01087; DB00794; DB01032; DB00396; DB00420; DB13602; DB09288; DB01182; DB12278; DB00571; DB06480; DB00545; DB01589; DB04216; DB01224; DB01103; DB13685; DB00908; DB00468; DB01369; DB12874; DB01129; DB00481; DB00980; DB00863; DB00243; DB00234; DB08896; DB11853; DB06458; DB14761; DB00409; DB00912; DB16826; DB02709; DB01256; DB13174; DB11730; DB06233; DB00615; DB04934; DB01045; DB11753; DB01201; DB01220; DB08864; DB12457; DB00896; DB06155; DB08931; DB14840; DB15305; DB00734; DB14924; DB00503; DB06228; DB09200; DB00533; DB01656; DB13409; DB09291; DB06176; DB00296; DB00412; DB05271; DB00778; DB12332; DB06201; DB11614; DB01698; DB08877; DB06654; DB12391; DB01001; DB00938; DB12543; DB01232; DB11805; DB11767; DB06335; DB00747; DB12834; DB14583; DB11459; DB01037; DB05885; DB11362; DB11942; DB15685; DB11689; DB06731; DB06739; DB06144; DB01104; DB01236; DB01105; DB00203; DB06207; DB09036; DB06290; DB00641; DB12371; DB00877; DB01261; DB06268; DB05482; DB01591; DB09308; DB09099; DB09143; DB00398; DB12713; DB15569; DB12548; DB01323; DB09118; DB00708; DB00359; DB01015; DB01138; DB01268; DB09034; DB09317; DB09318; DB00864; DB00820; DB00675; DB00706; DB06083; DB09071; DB01349; DB08833; DB12887; DB12020; DB05521; DB00976; DB12095; DB00231; DB06287; DB11761; DB00444; DB09299; DB15133; DB00857; DB00342; DB13399; DB13725; DB04905; DB00624; DB13943; DB13944; DB01420; DB13946; DB00759; DB12093; DB14066; DB11712; DB01041; DB00277; DB01154; DB00599; DB04572; DB00906; DB09289; DB08816; DB11470; DB00911; DB01007; DB01409; DB00932; DB06137; DB16732; DB11800; DB06273; DB11635; DB11251; DB08895; DB08811; DB09216; DB01036; DB06212; DB00273; DB01685; DB00539; DB05109; DB00193; DB08911; DB07615; DB00752; DB14962; DB05773; DB00656; DB00755; DB00620; DB00897; DB12245; DB12808; DB09089; DB00347; DB00440; DB06045; DB00197; DB13179; DB11652; DB15328; DB06267; DB08867; DB14989; DB13609; DB15091; DB01586; DB12255; DB11915; DB00580; DB00313; DB15114; DB05294; DB03701; DB04894; DB00862; DB11613; DB08881; DB11581; DB00285; DB00661; DB14895; DB06652; DB09082; DB06684; DB09185; DB00570; DB00541; DB00309; DB11641; DB00361; DB12131; DB08828; DB11094; DB00163; DB11693; DB11739; DB09030; DB00582; DB09068; DB14975; DB12026; DB00682; DB13950; DB01392; DB00549; DB00962; DB15035; DB15688; DB00495; DB00744; DB04832; DB00246; DB00425; DB04828; DB00909; DB01198; DB09225; DB01624; DB15490 Interacts with O15287; Q6ZQX7-4 EC number 1.14.14.1; 1.14.14.55; 1.14.14.56; 1.14.14.73 Uniprot keywords 3D-structure; Direct protein sequencing; Disease variant; Endoplasmic reticulum; Fatty acid metabolism; Heme; Iron; Lipid biosynthesis; Lipid metabolism; Membrane; Metal-binding; Microsome; Monooxygenase; Oxidoreductase; Proteomics identification; Reference proteome; Steroid biosynthesis; Steroid metabolism; Sterol metabolism; Transmembrane; Transmembrane helix; Ubl conjugation Protein physicochemical properties Chain ID A Molecular weight (Da) 52686.1 Length 460 Aromaticity 0.11 Instability index 43.51 Isoelectric point 8.62 Charge (pH=7) 4.97 3D Binding mode Sequence SHGLFKKLGIPGPTPLPFLGNILSYHKGFCMFDMECHKKYGKVWGFYDGQQPVLAITDPDMIKTVLVKECYSVFTNRRPFGPVGFMKSAISIAEDEEWKRLRSLLSPTFTSGKLKEMVPIIAQYGDVLVRNLRREAETGKPVTLKDVFGAYSMDVITSTSFGVNIDSLNNPQDPFVENTKKLLRFDFLDPFFLSITVFPFLIPILEVLNICVFPREVTNFLRKSVKRMKESRLEDTQKHRVDFLQLMIDSQALSDLELVAQSIIFIFAGYETTSSVLSFIMYELATHPDVQQKLQEEIDAVLPNKAPPTYDTVLQMEYLDMVVNETLRLFPIAMRLERVCKKDVEINGMFIPKGVVVMIPSYALHRDPKYWTEPEKFLPERFSKKNKDNIDPYIYTPFGSGPRNCIGMRFALMNMKLALIRVLQNFSFKPCKETQIPLKLSLGGLLQPEKPVVLKVESRD Hydrogen bonds contact Hydrophobic contact | ||||
| 13 | Tryptophan 5-hydroxylase 1 (TPH1) | 5TPG | 4.42 | |
Target general information Gen name TPH1 Organism Homo sapiens (Human) Uniprot ID TTD ID Synonyms Tryptophan 5-monooxygenase 1; TRPH; TPRH Protein family Biopterin-dependent aromatic amino acid hydroxylase family Biochemical class Paired donor oxygen oxidoreductase Function Responsible for addition of the -HO group (hydroxylation) to the 5 position to form the amino acid 5-hydroxytryptophan (5-HTP), which is the initial and rate-limiting step in the synthesis of the neurotransmitter serotonin. Related diseases Tyrosinemia 2 (TYRSN2) [MIM:276600]: An inborn error of metabolism characterized by elevations of tyrosine in the blood and urine, and oculocutaneous manifestations. Typical features include palmoplantar keratosis, painful corneal ulcers, and intellectual disability. {ECO:0000269|PubMed:1357662}. The disease is caused by variants affecting the gene represented in this entry. Drugs (DrugBank ID) DB05199; DB00360; DB12095; DB00150 Interacts with Q14457; Q96IK1-2; Q9UKB3; Q9H8Y8; O43586; O95789-4 EC number EC 1.14.16.4 Uniprot keywords 3D-structure; Alternative splicing; Iron; Metal-binding; Monooxygenase; Oxidoreductase; Phosphoprotein; Proteomics identification; Reference proteome; Serotonin biosynthesis; Ubl conjugation Protein physicochemical properties Chain ID A Molecular weight (Da) 31138.2 Length 271 Aromaticity 0.13 Instability index 43.43 Isoelectric point 6.73 Charge (pH=7) -0.86 3D Binding mode Sequence TVPWFPKKISDLDHCNVYRKRRKYFADLAMNYKHGDPIPKVEFTEEEIKTWGTVFQELNKLYPTHACREYLKNLPLLSKYCGYREDNIPQLEDVSNFLKERTGFSIRPVAGYLSPRDFLSGLAFRVFHCTQYVRHSSDPFYTPEPDTCHELLGHVPLLAEPSFAQFSQEIGLASLGASEEAVQKLATCYFFTVEFGLCKQDGQLRVFGAGLLSSISELKHALSGHAKVKPFDPKITCKQECLITTFQDVYFVSESFEDAKEKMREFTKTIK Hydrogen bonds contact Hydrophobic contact | ||||
| 14 | Urease subunit alpha | 1FWE | 4.42 | |
Target general information Gen name ureC Organism Klebsiella aerogenes (Enterobacter aerogenes) Uniprot ID TTD ID NA Synonyms NA Protein family Metallo-dependent hydrolases superfamily, Urease alpha subunit family Biochemical class Hydrolase Function Nickel cation binding.Urease activity. Related diseases Can contribute to cancer cell survival, proliferation, migration, and invasion, and tumor angiogenesis and metastasis. May contribute to cancer pathogenesis by promoting inflammatory responses and recruitment of tumor-infiltrating macrophages.; DISEASE: Abnormally high expression of soluble isoforms (isoform 2, isoform 3 or isoform 4) may be a cause of preeclampsia. Drugs (DrugBank ID) DB00551; DB05265 Interacts with P18316 EC number 3.5.1.5 Uniprot keywords 3D-structure; Cytoplasm; Hydrolase; Metal-binding; Nickel Protein physicochemical properties Chain ID C Molecular weight (Da) 80688.3 Length 753 Aromaticity 0.06 Instability index 28.4 Isoelectric point 5.58 Charge (pH=7) -21.17 3D Binding mode Sequence MELTPREKDKLLLFTAALVAERRLARGLKLNYPESVALISAFIMEGARDGKSVASLMEEGRHVLTREQVMEGVPEMIPDIQVEATFPDGSKLVTVHNPIISNISRQAYADMFGPTVGDKVRLADTELWIEVEDDLTTYGEEVKFGGGKVIRDGMGQGQMLAADCVDLVLTNALIVDHWGIVKADIGVKDGRIFAIGKAGNPDIQPNVTIPIGAATEVIAAEGKIVTAGGIDTHIHWICPQQAEEALVSGVTTMVGGGTGPAAGTHATTCTPGPWYISRMLQAADSLPVNIGLLGKGNVSQPDALREQVAAGVIGLXIHEDWGATPAAIDCALTVADEMDIQVALHSDTLNESGFVEDTLAAIGGRTIHTFHTEGAGGGHAPDIITACAHPNILPSSTNPTLPYTLNTIDEHLDMLMFAESRIRRETIAAEDVLHDLGAFSLTSSDSQAMGRVGEVILRTWQVAHRMKVQRGALAEETGDNDNFRVKRYIAKYTINPALTHGIAHEVGSIEVGKLADLVVWSPAFFGVKPATVIKGGMIAIAPMGDINASIPTPQPVHYRPMFGALGSARHHCRLTFLSQAAAANGVAERLNLRSAIAVVKGCRTVQKADMVHNSLQPNITVDAQTYEVRVDGELITSEPADVLPMAQRYFLFMIPGEYHVKPGQIALNTGRATCRVVVENHGDRPIQVGSHYHFAEVNPALKFDRQQAAGYRLNIPAGTAVRFEPGQKREVELVAFAGHRAVFGFRGEVMGPL Hydrogen bonds contact Hydrophobic contact | ||||
| 15 | Penicillin-binding protein 1A | 2ZC6 | 4.42 | |
Target general information Gen name ponA Organism Streptococcus pneumoniae serotype 4 (strain ATCC BAA-334 / TIGR4) Uniprot ID TTD ID NA Synonyms SP_0369 Protein family Glycosyltransferase 51 family; Transpeptidase family Biochemical class Biosynthetic protein Function Penicillin binding.Peptidoglycan glycosyltransferase activity.Serine-type D-Ala-D-Ala carboxypeptidase activity. Related diseases Aminoacylase-1 deficiency (ACY1D) [MIM:609924]: An enzymatic deficiency resulting in encephalopathy, unspecific psychomotor delay, psychomotor delay with atrophy of the vermis and syringomyelia, marked muscular hypotonia or normal clinical features. Epileptic seizures are a frequent feature. All affected individuals exhibit markedly increased urinary excretion of several N-acetylated amino acids. {ECO:0000269|PubMed:16274666, ECO:0000269|PubMed:16465618, ECO:0000269|PubMed:17562838, ECO:0000269|PubMed:21414403}. The disease is caused by variants affecting the gene represented in this entry. Drugs (DrugBank ID) DB01150; DB05659 Interacts with NA EC number 2.4.99.28; 3.4.16.4 Uniprot keywords 3D-structure; Antibiotic resistance; Carboxypeptidase; Cell shape; Cell wall biogenesis/degradation; Glycosyltransferase; Hydrolase; Multifunctional enzyme; Peptidoglycan synthesis; Protease; Reference proteome; Secreted; Transferase Protein physicochemical properties Chain ID A,C Molecular weight (Da) 44805.1 Length 400 Aromaticity 0.12 Instability index 31.82 Isoelectric point 4.88 Charge (pH=7) -14.68 3D Binding mode Sequence NYPAYMDNYLKEVINQVEEETGYNLLTTGMDVYTNVDQEAQKHLWDIYNTDEYVAYPDDELQVASTIVDVSNGKVIAQLGARHQSSNVSFGINQAVETNRDWGSTMKPITDYAPALEYGVYDSTATIVHDEPYNYPGTNTPVYNWDRGYFGNITLQYALQQSRNVPAVETLNKVGLNRAKTFLNGLGIDYPSIHYSNAISSNTTESDKKYGASSEKMAAAYAAFANGGTYYKPMYIHKVVFSDGSEKEFSNVGTRAMKETTAYMMTDMMKTVLSYGTGQNAYLAWLPQAGKTGTSNYTDEEIENHIKTSQFVAPDELFAGYTRKYSMAVWTGYSNRLTPLVGNGLTVAAKVYRSMMTYLSEGSNPEDWNIPEGLYRNGEFVFKNTSSKIYDNKNQLIADL Hydrogen bonds contact Hydrophobic contact | ||||
| 16 | N-acylethanolamine-hydrolyzing acidamidase (NAAA) | 6DXX | 4.42 | |
Target general information Gen name NAAA Organism Homo sapiens (Human) Uniprot ID TTD ID Synonyms Nacylsphingosine amidohydrolaselike; Nacylethanolaminehydrolyzing acid amidase subunit beta; NAAA; Acid ceramidaselike protein; ASAHlike protein Protein family Acid ceramidase family Biochemical class Carbon-nitrogen hydrolase Function Degrades bioactive fatty acid amides to their corresponding acids, with the following preference: N- palmitoylethanolamine > N-myristoylethanolamine > N- lauroylethanolamine = N-stearoylethanolamine > N- arachidonoylethanolamine > N-oleoylethanolamine. Also exhibits weak hydrolytic activity against the ceramides N- lauroylsphingosine and N-palmitoylsphingosine. Related diseases Hypertriglyceridemia, transient infantile (HTGTI) [MIM:614480]: An autosomal recessive disorder characterized by onset of moderate to severe transient hypertriglyceridemia in infancy that normalizes with age. The hypertriglyceridemia is associated with hepatomegaly, moderately elevated transaminases, persistent fatty liver, and the development of hepatic fibrosis. {ECO:0000269|PubMed:22226083, ECO:0000269|PubMed:24549054}. The disease is caused by variants affecting the gene represented in this entry. Drugs (DrugBank ID) DB09061; DB14009; DB14011 Interacts with NA EC number EC 3.5.1.- Uniprot keywords 3D-structure; Alternative splicing; Autocatalytic cleavage; Direct protein sequencing; Disulfide bond; Fatty acid metabolism; Glycoprotein; Hydrolase; Lipid degradation; Lipid metabolism; Lysosome; Membrane; Proteomics identification; Reference proteome; Signal; Zymogen Protein physicochemical properties Chain ID A,B Molecular weight (Da) 36877.8 Length 328 Aromaticity 0.11 Instability index 44.37 Isoelectric point 7.72 Charge (pH=7) 1.08 3D Binding mode Sequence SPPAAPRFNVSLDSVPELRWLPVLRHYDLDLVRAAMAQVIGDRVPKWVHVLIGKVVLELERFLPQPFTGEIRGMCDFMNLSLADCLLVNLAYESSVFCTSIVAQDSRGHIYHGRNLDYPFGNVLRKLTVDVQFLKNGQIAFTGTTFIGYVGLWTGQSPHKFTVSGDERDKGWWWENAIAALFRRHIPVSWLIRATLSESENFEAAVGKLAKTPLIADVYYIVGGTSPREGVVITRNRDGPADIWPLDPLNGAWFRVETNYDHWKPAPKEDDRRTSAIKALNATGQANLSLEALFQILSVVPVYNNFTIYTTVMSAGSPDKYMTRIRNP Hydrogen bonds contact Hydrophobic contact | ||||
| 17 | C-C chemokine receptor type 2 (CCR2) | 5T1A | 4.42 | |
Target general information Gen name CCR2 Organism Homo sapiens (Human) Uniprot ID TTD ID Synonyms Monocyte chemoattractant protein 1 receptor; MCP-1-R; Chemokine receptor CCR2B; CMKBR2; CD192; CCR-2; CC-CKR-2; C-C CKR-2 Protein family G-protein coupled receptor 1 family Biochemical class GPCR rhodopsin Function Its binding with CCL2 on monocytes and macrophages mediates chemotaxis and migration induction through the activation of the PI3K cascade, the small G protein Rac and lamellipodium protrusion. Also acts as a receptor for the beta-defensin DEFB106A/DEFB106B. Regulates the expression of T-cell inflammatory cytokines and T-cell differentiation, promoting the differentiation of T-cells into T-helper 17 cells (Th17) during inflammation. Faciltates the export of mature thymocytes by enhancing directional movement of thymocytes to sphingosine-1-phosphate stimulation and up-regulation of S1P1R expression; signals through the JAK-STAT pathway to regulate FOXO1 activity leading to an increased expression of S1P1R. Plays an important role in mediating peripheral nerve injury-induced neuropathic pain. Increases NMDA-mediated synaptic transmission in both dopamine D1 and D2 receptor-containing neurons, which may be caused by MAPK/ERK-dependent phosphorylation of GRIN2B/NMDAR2B. Mediates the recruitment of macrophages and monocytes to the injury site following brain injury. Key functional receptor for CCL2 but can also bind CCL7 and CCL12. Related diseases Polycystic lung disease (PCLUD) [MIM:219600]: An autosomal recessive disease characterized by pulmonary alveolar proteinosis, marked peribronchovascular and parenchymal lymphocytosis, peribronchiolar pulmonary fibrosis, progressive diffuse parenchymal lung cyst formation and enlargement, progressive obstructive airflow limitation, and recurrent secondary infections. Additional features may include digital clubbing, allergies, and atopic dermatitis. {ECO:0000269|PubMed:38157855}. The disease is caused by variants affecting the gene represented in this entry. Drugs (DrugBank ID) DB05159; DB11758; DB05130; DB12520 Interacts with Q6S8J3; Q9BW27 EC number NA Uniprot keywords 3D-structure; Alternative splicing; Cell membrane; Disease variant; Disulfide bond; G-protein coupled receptor; Glycoprotein; Host-virus interaction; Inflammatory response; Membrane; Phosphoprotein; Proteomics identification; Receptor; Reference proteome; Sulfation; Transducer; Transmembrane; Transmembrane helix Protein physicochemical properties Chain ID A Molecular weight (Da) 50270.6 Length 445 Aromaticity 0.13 Instability index 29.96 Isoelectric point 9.49 Charge (pH=7) 17.19 3D Binding mode Sequence VKQIGAQLLPPLYSLVFIFGFVGNMLVVLILINCKKLKCLTDIYLLNLAISDLLFLITLPLWAHSAANEWVFGNAMCKLFTGLYHIGYFGGIFFIILLTIDRYLAIVHAVFALKARTVTFGVVTSVITWLVAVFASVPGIIFTKXQKEDSVYVCGPYFPRGWNNFHTIMRNILGLVLPLLIMVICYSGISRASKSRINIFEMLRIDEGLRLKIYKDTEGYYTIGIGHLLTKSPSLNAAKSELDKAIGRNTNGVITKDEAEKLFNQDVDAAVRGILRNAKLKPVYDSLDAVRRAALINMVFQMGETGVAGFTNSLRMLQQKRWDEAAVNLAKSRWYNQTPNRAKRVITTFRTGTWDAYPPPSREKKAVRVIFTIMIVYFLFWTPYNIVILLNTFQEFFGLSNCESTSQLDQATQVTETLGMTHCCINPIIYAFVGEKFRRYLSVFF Hydrogen bonds contact Hydrophobic contact | ||||
| 18 | Histone deacetylase 7 (HDAC7) | 3C0Z | 4.42 | |
Target general information Gen name HDAC7 Organism Homo sapiens (Human) Uniprot ID TTD ID Synonyms Histone deacetylase 7A; HDAC7A; HD7a; HD7 Protein family Histone deacetylase family, HD type 2 subfamily Biochemical class Carbon-nitrogen hydrolase Function Gives a tag for epigenetic repression and plays an important role in transcriptional regulation, cell cycle progression and developmental events. Histone deacetylases act via the formation of large multiprotein complexes. Involved in muscle maturation by repressing transcription of myocyte enhancer factors such as MEF2A, MEF2B and MEF2C. During muscle differentiation, it shuttles into the cytoplasm, allowing the expression of myocyte enhancer factors. May be involved in Epstein-Barr virus (EBV) latency, possibly by repressing the viral BZLF1 gene. Positively regulates the transcriptional repressor activity of FOXP3. Responsible for the deacetylation of lysine residues on the N-terminal part of the core histones (H2A, H2B, H3 and H4). Related diseases Mucopolysaccharidosis 1H (MPS1H) [MIM:607014]: A severe form of mucopolysaccharidosis type 1, a rare lysosomal storage disease characterized by progressive physical deterioration with urinary excretion of dermatan sulfate and heparan sulfate. Patients with MPS1H usually present, within the first year of life, a combination of hepatosplenomegaly, skeletal deformities, corneal clouding and severe intellectual disability. Obstructive airways disease, respiratory infection and cardiac complications usually result in death before 10 years of age. {ECO:0000269|PubMed:10466419, ECO:0000269|PubMed:10735634, ECO:0000269|PubMed:12559846, ECO:0000269|PubMed:1301941, ECO:0000269|PubMed:15300847, ECO:0000269|PubMed:19396826, ECO:0000269|PubMed:21394825, ECO:0000269|PubMed:24036510, ECO:0000269|PubMed:31194252, ECO:0000269|PubMed:7550232, ECO:0000269|PubMed:7550242, ECO:0000269|PubMed:7951228, ECO:0000269|PubMed:8019563, ECO:0000269|PubMed:8328452, ECO:0000269|PubMed:8401515, ECO:0000269|Ref.20}. The disease is caused by variants affecting the gene represented in this entry.; DISEASE: Mucopolysaccharidosis 1H/S (MPS1H/S) [MIM:607015]: A form of mucopolysaccharidosis type 1, a rare lysosomal storage disease characterized by progressive physical deterioration with urinary excretion of dermatan sulfate and heparan sulfate. MPS1H/S represents an intermediate phenotype of the MPS1 clinical spectrum. It is characterized by relatively little neurological involvement, but most of the somatic symptoms described for severe MPS1 develop in the early to mid-teens, causing considerable loss of mobility. {ECO:0000269|PubMed:10466419, ECO:0000269|PubMed:10735634, ECO:0000269|PubMed:12559846, ECO:0000269|PubMed:15300847, ECO:0000269|PubMed:21394825, ECO:0000269|PubMed:7550232, ECO:0000269|PubMed:7550242, ECO:0000269|PubMed:8401515}. The disease is caused by variants affecting the gene represented in this entry.; DISEASE: Mucopolysaccharidosis 1S (MPS1S) [MIM:607016]: A mild form of mucopolysaccharidosis type 1, a rare lysosomal storage disease characterized by progressive physical deterioration with urinary excretion of dermatan sulfate and heparan sulfate. Patients with MPS1S may have little or no neurological involvement, normal stature and life span, but present development of joints stiffness, mild hepatosplenomegaly, aortic valve disease and corneal clouding. {ECO:0000269|PubMed:12559846, ECO:0000269|PubMed:15300847, ECO:0000269|PubMed:19396826, ECO:0000269|PubMed:21394825, ECO:0000269|PubMed:25256405, ECO:0000269|PubMed:7550232, ECO:0000269|PubMed:7550242, ECO:0000269|PubMed:8213840}. The disease is caused by variants affecting the gene represented in this entry. Drugs (DrugBank ID) DB12565; DB05015; DB01262; DB11841; DB12645; DB06603; DB06819; DB03766; DB12847; DB06176; DB04297; DB00313; DB02546 Interacts with P00533; Q9BZS1-1; Q9BZS1-2; Q9BZL6; P31947; P63104; P08393; Q8CFN5; Q13137; Q04864; Q0D2K3; Q8WXI4-2; Q9BQD7; Q03989; Q9NSI6-4; Q3SXR2; Q13137; P60953; Q7L2Z9; Q96D03; Q9NQ30; Q9UBI6; A6NEM1; A5PKX9; Q9BXK1; Q6ZNG9; O43679; Q6FHY5; Q9BRT3; O94964-4; O95411; O00746; Q9BQI9; B7ZLY0; Q96I34; P63000; P15153; P60763; Q04864-2; Q0D2K3; P62070; O15427; O95164 EC number EC 3.5.1.98 Uniprot keywords 3D-structure; Alternative splicing; Chromatin regulator; Cytoplasm; Hydrolase; Metal-binding; Nucleus; Phosphoprotein; Proteomics identification; Reference proteome; Repeat; Repressor; Transcription; Transcription regulation; Zinc Protein physicochemical properties Chain ID A Molecular weight (Da) 41454.5 Length 383 Aromaticity 0.08 Instability index 38.49 Isoelectric point 6.26 Charge (pH=7) -5.18 3D Binding mode Sequence TLPFTTGLIYDSVMLKHQCSCGDNSRHPEHAGRIQSIWSRLQERGLRSQCECLRGRKASLEELQSVHSERHVLLYGTNPLSRLKLDNGKLAGLLAQVMLPCGGVGVDTDTIWNELHSSNAARWAAGSVTDLAFKVASRELKNGFAVVRPPGHHADHSTAMGFCFFNSVAIACRQLQQQSKASKILIVDWDVHHGNGTQQTFYQDPSVLYISLHRHDDGNFFPGSGAVDEVGAGSGEGFNVNVAWAGGLDPPMGDPEYLAAFRIVVMPIAREFSPDLVLVSAGFDAAEGHPAPLGGYHVSAKCFGYMTQQLMNLAGGAVVLALEGGHDLTAICDASEACVAALLGNRVDPLSEEGWKQKPNLNAIRSLEAVIRVHSKYWGCMQR Hydrogen bonds contact Hydrophobic contact | ||||
| 19 | Gamma-aminobutyric acid type B receptor subunit 1 | 4MS4 | 4.41 | |
Target general information Gen name GABBR1 Organism Homo sapiens (Human) Uniprot ID TTD ID NA Synonyms GPRC3B;GPR51 Protein family G-protein coupled receptor 3 family, GABA-B receptor subfamily Biochemical class Signaling protein / antagonist Function G-protein coupled GABA receptor activity. Related diseases Neurodevelopmental disorder with poor language and loss of hand skills (NDPLHS) [MIM:617903]: An autosomal dominant disorder characterized by psychomotor developmental stagnation or regression. NDPLHS manifest in the first years of life as loss of purposeful hand movements, loss of language, and intellectual disability. {ECO:0000269|PubMed:26740508, ECO:0000269|PubMed:28856709, ECO:0000269|PubMed:29369404}. The disease is caused by variants affecting the gene represented in this entry.; DISEASE: Developmental and epileptic encephalopathy 59 (DEE59) [MIM:617904]: A form of epileptic encephalopathy, a heterogeneous group of severe early-onset epilepsies characterized by refractory seizures, neurodevelopmental impairment, and poor prognosis. Development is normal prior to seizure onset, after which cognitive and motor delays become apparent. DEE59 is an autosomal dominant condition characterized by onset of refractory seizures in early infancy. {ECO:0000269|PubMed:28856709, ECO:0000269|PubMed:29100083, ECO:0000269|PubMed:29369404}. The disease is caused by variants affecting the gene represented in this entry. Drugs (DrugBank ID) DB08891; DB08892; DB00181; DB00363; DB02530; DB05010; DB09072 Interacts with Q9UBS5; Q9UBS5-2; P46459; Q86UR5 EC number NA Uniprot keywords 3D-structure; Cell membrane; Coiled coil; Direct protein sequencing; Disease variant; Disulfide bond; Epilepsy; G-protein coupled receptor; Glycoprotein; Intellectual disability; Membrane; Phosphoprotein; Postsynaptic cell membrane; Proteomics identification; Receptor; Reference proteome; Signal; Synapse; Transducer; Transmembrane; Transmembrane helix Protein physicochemical properties Chain ID A Molecular weight (Da) 46502.1 Length 408 Aromaticity 0.12 Instability index 50.05 Isoelectric point 5.78 Charge (pH=7) -5.62 3D Binding mode Sequence RRAVYIGALFPMSGGWPGGQACQPAVEMALEDVNSRRDILPDYELKLIHHDSKCDPGQATKYLYELLYNDPIKIILMPGCSSVSTLVAEAARMWNLIVLSYGSSSPALSNRQRFPTFFRTHPSATLHNPTRVKLFEKWGWKKIATIQQTTEVFTSTLDDLEERVKEAGIEITFRQSFFSDPAVPVKNLKRQDARIIVGLFYETEARKVFCEVYKERLFGKKYVWFLIGWYADNWFKIYDPSINCTVDEMTEAVEGHITTEIVMLNPANTRSISNMTSQEFVEKLTKRLKRHPEETGGFQEAPLAYDAIWALALALNKTSRLEDFNYNNQTITDQIYRAMNSSSFEGVSGHVVFDASGSRMAWTLIEQLQGGSYKKIGYYDSTKDDLSWSKTDKWIGGSPPADDYKDDD Hydrogen bonds contact Hydrophobic contact | ||||
| 20 | mRNA-capping enzyme | 2C46 | 4.41 | |
Target general information Gen name RNGTT Organism Homo sapiens (Human) Uniprot ID TTD ID NA Synonyms CAP1A Protein family Non-receptor class of the protein-tyrosine phosphatase family; Eukaryotic GTase family Biochemical class Transferase Function GTP binding.MRNA guanylyltransferase activity.Polynucleotide 5'-phosphatase activity.Protein tyrosine/serine/threonine phosphatase activity.Protein tyrosine phosphatase activity.RNA guanylyltransferase activity.Triphosphatase activity. Related diseases Atrial fibrillation, familial, 14 (ATFB14) [MIM:615378]: A familial form of atrial fibrillation, a common sustained cardiac rhythm disturbance. Atrial fibrillation is characterized by disorganized atrial electrical activity and ineffective atrial contraction promoting blood stasis in the atria and reduces ventricular filling. It can result in palpitations, syncope, thromboembolic stroke, and congestive heart failure. {ECO:0000269|PubMed:19808477}. The disease is caused by variants affecting the gene represented in this entry.; DISEASE: Genetic variations in SCN2B may be involved in Brugada syndrome (PubMed:23559163). This tachyarrhythmia is characterized by right bundle branch block and ST segment elevation on an electrocardiogram (ECG). It can cause the ventricles to beat so fast that the blood is prevented from circulating efficiently in the body. When this situation occurs, the individual will faint and may die in a few minutes if the heart is not reset. {ECO:0000269|PubMed:23559163}. Drugs (DrugBank ID) NA Interacts with Q92624; P16333-1 EC number 2.7.7.50; 3.6.1.74 Uniprot keywords 3D-structure; Alternative splicing; GTP-binding; Host-virus interaction; Hydrolase; mRNA capping; mRNA processing; Multifunctional enzyme; Nucleotide-binding; Nucleotidyltransferase; Nucleus; Protein phosphatase; Proteomics identification; Reference proteome; Transferase Protein physicochemical properties Chain ID A,B,C,D Molecular weight (Da) 21849.8 Length 189 Aromaticity 0.11 Instability index 53.71 Isoelectric point 5.89 Charge (pH=7) -2.91 3D Binding mode Sequence NKIPPRWLNCPRRGQPVAGRFLPLKTMLGPRYDSQVAEENRFHPSMLSNYLKSVKMGLLVDLTNTSRFYDRNDIEKEGIKYIKLQCKGHGECPTTENTETFIRLCERFELIGVHCTHGFNRTGFLICAFLVEKMDWSIEAAVATFAQARPPGIYKGDYLKELFRRYGDIEEAPPPPLLPDWCFEDDEDE Hydrogen bonds contact Hydrophobic contact | ||||