Job Results:
Ligand
Structure
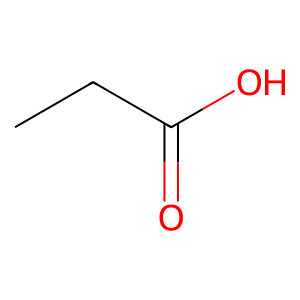
Job ID
1aab99f222a82120b630ad6d44e2ad5f
Job name
NA
Time
2026-02-27 11:58:02
| Rank | Target | PDB ID |
AirScore |
Detail |
|---|---|---|---|---|
| 1 | Transferrin (TF) | 1RYO | 4.66 | |
Target general information Gen name TF Organism Homo sapiens (Human) Uniprot ID TTD ID Synonyms Siderophilin; Serotransferrin; PRO1400; Beta-1 metal-binding globulin Protein family Transferrin family Biochemical class Transferrin Function It is responsible for the transport of iron from sites of absorption and heme degradation to those of storage and utilization. Serum transferrin may also have a further role in stimulating cell proliferation. Transferrins are iron binding transport proteins which can bind two Fe(3+) ions in association with the binding of an anion, usually bicarbonate. Related diseases Atransferrinemia (ATRAF) [MIM:209300]: A rare autosomal recessive disorder characterized by abnormal synthesis of transferrin leading to iron overload and microcytic hypochromic anemia. {ECO:0000269|PubMed:11110675, ECO:0000269|PubMed:15466165}. The disease is caused by variants affecting the gene represented in this entry. Drugs (DrugBank ID) DB01370; DB14517; DB14518; DB01294; DB14526; DB14527; DB11136; DB14528; DB14529; DB14530; DB00515; DB09130; DB11397; DB13949; DB14490; DB14491; DB14488; DB14501; DB14489; DB13257; DB06215; DB06784; DB05260; DB01592; DB00893; DB00677; DB06757; DB11182; DB14520; DB01593; DB14487; DB14533; DB14548 Interacts with O43315; O00501; Q7Z7G2; Q9GZR5; Q9Y282; Q96KR6; P01350; P08034; Q8NBJ4; O15529; Q8TED1; Q7Z5P4; A8MZ59; O15173; Q96TC7; Q3KNW5; Q9BXS9-3; Q99523; O43278-2; Q8N9I0; P02786; Q4KMG9; Q9K0U9; Q09057; Q9K0V0; P02786 EC number NA Uniprot keywords 3D-structure; Direct protein sequencing; Disease variant; Disulfide bond; Glycoprotein; Ion transport; Iron; Iron transport; Metal-binding; Methylation; Phosphoprotein; Proteomics identification; Reference proteome; Repeat; Secreted; Signal; Transport Protein physicochemical properties Chain ID A Molecular weight (Da) 35854.5 Length 324 Aromaticity 0.1 Instability index 34.17 Isoelectric point 7.58 Charge (pH=7) 1.42 3D Binding mode Sequence KTVRWCAVSEHEATKCQSFRDHMKSVIPSDGPSVACVKKASYLDCIRAIAANEADAVTLDAGLVYDAYLAPNNLKPVVAEFYGSKEDPQTFYYAVAVVKKDSGFQMNQLRGKKSCHTGLGRSAGWNIPIGLLYCDLPEPRKPLEKAVANFFSGSCAPCADGTDFPQLCQLCPGCGCSTLNQYFGYSGAFKCLKDGAGDVAFVKHSTIFENLANKADRDQYELLCLDNTRKPVDEYKDCHLAQVPSHTVVARSMGGKEDLIWELLNQAQEHFGKDKSKEFQLFSSPHGKDLLFKDSAHGFLKVPPRMDAKMYLGYEYVTAIRNLR Hydrogen bonds contact Hydrophobic contact | ||||
| 2 | Ubiquitin carboxyl-terminal hydrolase 2 (USP2) | 5XU8 | 4.66 | |
Target general information Gen name USP2 Organism Homo sapiens (Human) Uniprot ID TTD ID Synonyms Ubiquitin-specific-processing protease 2; Ubiquitin thioesterase 2; UBP41; Deubiquitinating enzyme 2; 41 kDa ubiquitin-specific protease Protein family Peptidase C19 family, USP2 subfamily Biochemical class Peptidase Function Isoform 1 and isoform 4 possess both ubiquitin-specific peptidase and isopeptidase activities. Deubiquitinates MDM2 without reversing MDM2-mediated p53/TP53 ubiquitination and thus indirectly promotes p53/TP53 degradation and limits p53 activity. Has no deubiquitinase activity against p53/TP53. Prevents MDM2-mediated degradation of MDM4. Plays a role in the G1/S cell-cycle progression in normal and cancer cells. Regulates the circadian clock by modulating its intrinsic circadian rhythm and its capacity to respond to external cues. Associates with clock proteins and deubiquitinates core clock component PER1 but does not affect its overall stability. Regulates the nucleocytoplasmic shuttling and nuclear retention of PER1 and its repressive role on the clock transcription factors CLOCK and ARNTL/BMAL1. Plays a role in the regulation of myogenic differentiation of embryonic muscle cells. Hydrolase that deubiquitinates polyubiquitinated target proteins such as MDM2, MDM4 and CCND1. Related diseases Defects in AKT2 are a cause of susceptibility to breast cancer (BC). AKT2 promotes metastasis of tumor cells without affecting the latency of tumor development. May play a role in glioblastoma cell survival (PubMed:20167810). {ECO:0000269|PubMed:20167810}.; DISEASE: Type 2 diabetes mellitus (T2D) [MIM:125853]: A multifactorial disorder of glucose homeostasis caused by a lack of sensitivity to insulin. Affected individuals usually have an obese body habitus and manifestations of a metabolic syndrome characterized by diabetes, insulin resistance, hypertension and hypertriglyceridemia. The disease results in long-term complications that affect the eyes, kidneys, nerves, and blood vessels. {ECO:0000269|PubMed:15166380, ECO:0000269|PubMed:19164855}. Disease susceptibility is associated with variants affecting the gene represented in this entry.; DISEASE: Hypoinsulinemic hypoglycemia with hemihypertrophy (HIHGHH) [MIM:240900]: A disorder characterized by hypoglycemia, low insulin levels, low serum levels of ketone bodies and branched-chain amino acids, left-sided hemihypertrophy, neonatal macrosomia, reduced consciousness and hypoglycemic seizures. {ECO:0000269|PubMed:21979934}. The disease is caused by variants affecting the gene represented in this entry. Drugs (DrugBank ID) NA Interacts with Q9NYB9-2; P12814; P35609; Q08043; Q86U10; Q86V38; P56945; Q8TD16-2; Q96CA5; A2RRN7; Q13137; Q9H257-2; Q96JN2-2; Q2TAC2; A6NC98; Q96MT8-3; Q8NHQ1; Q9BSW2; Q8N4Y2-3; Q8WTU0; O75140-2; Q9NRI5-2; Q8N9I9; Q9H596; Q8WWB3; Q5JST6; Q9NRA8; O00471; Q96B26; P57678; Q08379; Q9NYA3; A6NEM1; Q6PI77; Q14451-3; Q4V328; Q9NSC5; Q9UJC3; Q96ED9-2; Q8IYA8; Q9UKT9; Q5TA45; Q96N16; O75564-2; Q674X7-2; Q9BVG8; Q9BVG8-5; P19012; Q7Z3Y8; Q15323; Q14525; O76011; Q92764; Q6A162; Q9UBR4-2; Q969G2; Q03252; Q9BRK4; Q00987; Q9UJV3-2; Q5VZ52; Q13084; Q5JR59; Q5JR59-3; Q15742; Q9GZM8; I6L9F6; P07196; O43482; Q96CV9; Q4G0R1; Q9NRD5; Q58EX7; Q8ND90; Q16633; Q9GZV8; Q6MZQ0; Q15276; Q8HWS3; Q59EK9-3; P60903; O14492-2; O60504; Q99932-2; A6NLX3; P51692; Q86VP1; Q8WW24; Q9UBB9; Q08117-2; Q03169; Q13077; Q12933; Q9Y4K3; P36406; P14373; Q86XT4; Q15654; Q8N6Y0; Q70EL1-9; Q9UK41-2; Q8N1B4; O96006; Q9NZV7; Q9UGI0; P05067; P54253; G5E9A7; Q01658; Q00403; Q9Y5Q9; P04792; O43464; P42858; Q8WXH2; O60333-2; A0A6Q8PF08; O60260-5; P60891; Q9Y3C5; Q7Z333; P37840; P00441; Q7Z699; Q13148; O76024 EC number EC 3.4.19.12 Uniprot keywords 3D-structure; Alternative splicing; Biological rhythms; Cell cycle; Cytoplasm; Hydrolase; Membrane; Metal-binding; Myogenesis; Nucleus; Protease; Proteomics identification; Reference proteome; Thiol protease; Ubl conjugation pathway; Zinc Protein physicochemical properties Chain ID A Molecular weight (Da) 37785.5 Length 327 Aromaticity 0.11 Instability index 42.45 Isoelectric point 8.23 Charge (pH=7) 3.56 3D Binding mode Sequence QGLAGLRNLGNTCFMNSILQCLSNTRELRDYCLQRLYMRDLHHGSNAHTALVEEFAKLIQTIWTSSPNDVVSPSEFKTQIQRYAPRFVGYNQQDAQEFLRFLLDGLHNEVNRVNLDHLPDDEKGRQMWRKYLEREDSRIGDLFVGQLKSSLTCTDCGYCSTVFDPFWDLSLPIAKRGYPEVTLMDCMRLFTKEDVLDGDEKPTCCRCRGRKRCIKKFSIQRFPKILVLHLKRFSESRIRTSKLTTFVNFPLRDLDLREFASENTNHAVYNLYAVSNHSGTTMGGHYTAYCRSPGTGEWHTFNDSSVTPMSSSQVRTSDAYLLFYELA Hydrogen bonds contact Hydrophobic contact | ||||
| 3 | Cystathionine gamma-lyase (CTH) | 3COG | 4.65 | |
Target general information Gen name CTH Organism Homo sapiens (Human) Uniprot ID TTD ID Synonyms Gamma-cystathionase; Cysteine-protein sulfhydrase Protein family Trans-sulfuration enzymes family Biochemical class NA Function Catalyzes the last step in the trans-sulfuration pathway from methionine to cysteine. Has broad substrate specificity. Converts cystathionine to cysteine, ammonia and 2-oxobutanoate. Converts two cysteine molecules to lanthionine and hydrogen sulfide. Can also accept homocysteine as substrate. Specificity depends on the levels of the endogenous substrates. Generates the endogenous signaling molecule hydrogen sulfide (H2S), and so contributes to the regulation of blood pressure. Acts as a cysteine-protein sulfhydrase by mediating sulfhydration of target proteins: sulfhydration consists of converting -SH groups into -SSH on specific cysteine residues of target proteins such as GAPDH, PTPN1 and NF-kappa-B subunit RELA, thereby regulating their function. Related diseases Cystathioninuria (CSTNU) [MIM:219500]: Autosomal recessive phenotype characterized by abnormal accumulation of plasma cystathionine, leading to increased urinary excretion. {ECO:0000269|PubMed:12574942, ECO:0000269|PubMed:18476726}. The disease is caused by variants affecting the gene represented in this entry. Drugs (DrugBank ID) DB02328; DB03928; DB00151; DB04217; DB00114 Interacts with P32929; Q96NT3; Q96NT3-2; Q96HA8; Q6P9E2 EC number EC 4.4.1.1 Uniprot keywords 3D-structure; Alternative splicing; Amino-acid biosynthesis; Calmodulin-binding; Cysteine biosynthesis; Cytoplasm; Disease variant; Lipid metabolism; Lyase; Proteomics identification; Pyridoxal phosphate; Reference proteome Protein physicochemical properties Chain ID A,B,C,D Molecular weight (Da) 86026 Length 782 Aromaticity 0.08 Instability index 32.4 Isoelectric point 6.27 Charge (pH=7) -9.46 3D Binding mode Sequence GFLPHFQHFATQAIHVGQDPEQWTSRAVVPPISLSTTFKQGAPGQHSGFEYSRSGNPTRNCLEKAVAALDGAKYCLAFASGLAATVTITHLLKAGDQIICMDDVYGGTNRYFRQVASEFGLKISFVDCSKIKLLEAAITPETKLVWIETPTNPTQKVIDIEGCAHIVHKHGDIILVVDNTFMSPYFQRPLALGADISMYSATKYMNGHSDVVMGLVSVNCESLHNRLRFLQNSLGAVPSPIDCYLCNRGLKTLHVRMEKHFKNGMAVAQFLESNPWVEKVIYPGLPSHPQHELVKRQCTGCTGMVTFYIKGTLQHAEIFLKNLKLFTLAESLGGFESLAELPAIMTHASVLKNDRDVLGISDTLIRLSVGLEDEEDLLEDLDQALKAAHPPSGFLPHFQHFATQAIHVGQDPEQWTSRAVVPPISLSTTFKQGAPGQGFEYSRSGNPTRNCLEKAVAALDGAKYCLAFASGLAATVTITHLLKAGDQIICMDDVYGGTNRYFRQVASEFGLKISFVDCSKIKLLEAAITPETKLVWIETPTNPTQKVIDIEGCAHIVHKHGDIILVVDNTFMSPYFQRPLALGADISMYSATKYMNGHSDVVMGLVSVNCESLHNRLRFLQNSLGAVPSPIDCYLCNRGLKTLHVRMEKHFKNGMAVAQFLESNPWVEKVIYPGLPSHPQHELVKRQCTGCTGMVTFYIKGTLQHAEIFLKNLKLFTLAESLGGFESLAELPAIMTHASVLKNDRDVLGISDTLIRLSVGLEDEEDLLEDLDQALKAAHPPS Hydrogen bonds contact Hydrophobic contact | ||||
| 4 | Endolysin | 1AM7 | 4.64 | |
Target general information Gen name R Organism Escherichia phage lambda (Bacteriophage lambda) Uniprot ID TTD ID NA Synonyms NA Protein family Glycosyl hydrolase 24 family Biochemical class Glycosidase Function Lyase activity.Lysozyme activity.Lytic transglycosylase activity. Related diseases Estrogen resistance (ESTRR) [MIM:615363]: A disorder characterized by partial or complete resistance to estrogens, in the presence of elevated estrogen serum levels. Clinical features include absence of the pubertal growth spurt, delayed bone maturation, unfused epiphyses, reduced bone mineral density, osteoporosis, continued growth into adulthood and very tall adult stature. Glucose intolerance, hyperinsulinemia and lipid abnormalities may also be present. {ECO:0000269|PubMed:23841731, ECO:0000269|PubMed:27754803}. The disease is caused by variants affecting the gene represented in this entry. Drugs (DrugBank ID) DB04206 Interacts with NA EC number 4.2.2.n2 Uniprot keywords 3D-structure; Antimicrobial; Bacteriolytic enzyme; Cytolysis; Direct protein sequencing; Host cell lysis by virus; Host cytoplasm; Lyase; Reference proteome; Viral release from host cell Protein physicochemical properties Chain ID A,B,C Molecular weight (Da) 49834.9 Length 462 Aromaticity 0.07 Instability index 18.78 Isoelectric point 9.6 Charge (pH=7) 18.29 3D Binding mode Sequence MVEINNQRKAFLDMLAXSEGTDNGRQKTRNHGYDVIVGGELFTDYSDHPRKLVTLNPKLKSTGAGRYQLLSRXXDAYRKQLGLKDFSPKSQDAVALQQIKERGALPMIDRGDIRQAIDRCSNIXASLPGAGYGQFEHKADSLIAKFKEAGGTVRMVEINNQRKAFLDMLAXSEGTDNGRQKTRNHGYDVIVGGELFTDYSDHPRKLVTLNPKLKSTGAGRYQLLSRXXDAYRKQLGLKDFSPKSQDAVALQQIKERGALPMIDRGDIRQAIDRCSNIXASLPGAGYGQFEHKADSLIAKFKEAGGTVRMVEINNQRKAFLDMLAXSEGTDNGRQKTRNHGYDVIVGGELFTDYSDHPRKLVTLNPKLKSTGAGRYQLLSRXXDAYRKQLGLKDFSPKSQDAVALQQIKERGALPMIDRGDIRQAIDRCSNIXASLPGAGYGQFEHKADSLIAKFKEAGGTVR Hydrogen bonds contact Hydrophobic contact | ||||
| 5 | Peptidyl-prolyl cis-trans isomerase G | 2GW2 | 4.64 | |
Target general information Gen name PPIG Organism Homo sapiens (Human) Uniprot ID TTD ID NA Synonyms NA Protein family NA Biochemical class Isomerase Function Cyclosporin A binding.Peptidyl-prolyl cis-trans isomerase activity.RNA binding. Related diseases Intellectual developmental disorder, autosomal dominant 6, with or without seizures (MRD6) [MIM:613970]: A disorder characterized by significantly below average general intellectual functioning associated with impairments in adaptive behavior and manifested during the developmental period. MRD6 additional features may include seizures, hypotonia, abnormal movements, such as dystonia, and autistic features. {ECO:0000269|PubMed:20890276, ECO:0000269|PubMed:23033978, ECO:0000269|PubMed:23160955, ECO:0000269|PubMed:24863970, ECO:0000269|PubMed:25356899, ECO:0000269|PubMed:27839871, ECO:0000269|PubMed:28095420, ECO:0000269|PubMed:38538865}. The disease is caused by variants affecting the gene represented in this entry.; DISEASE: Developmental and epileptic encephalopathy 27 (DEE27) [MIM:616139]: A form of epileptic encephalopathy, a heterogeneous group of severe early-onset epilepsies characterized by refractory seizures, neurodevelopmental impairment, and poor prognosis. Development is normal prior to seizure onset, after which cognitive and motor delays become apparent. {ECO:0000269|PubMed:24272827, ECO:0000269|PubMed:27839871, ECO:0000269|PubMed:27864847, ECO:0000269|PubMed:38538865}. The disease is caused by variants affecting the gene represented in this entry.; DISEASE: A chromosomal aberrations involving GRIN2B has been found in patients with intellectual disability. Translocations t(9;12)(p23;p13.1) and t(10;12)(q21.1;p13.1) with a common breakpoint in 12p13.1. Drugs (DrugBank ID) DB00172 Interacts with Q8N7W2-2; Q8NHQ1; O75553; Q9UI36-2; Q96C98; Q8NC69; P17931; Q6NVH9; Q15365; Q9UL42; Q96CD2; Q14498; Q16637; Q12800; Q9NVV9; PRO_0000037309 [P0C6X7] EC number 5.2.1.8 Uniprot keywords 3D-structure; Alternative splicing; Isomerase; Isopeptide bond; Nucleus; Phosphoprotein; Proteomics identification; Reference proteome; Rotamase; Ubl conjugation Protein physicochemical properties Chain ID A Molecular weight (Da) 19125.4 Length 173 Aromaticity 0.1 Instability index 26.46 Isoelectric point 7.14 Charge (pH=7) 0.24 3D Binding mode Sequence RPRCFFDIAINNQPAGRVVFELFSDVCPKTCENFRCLCTGEKGTGKSTQKPLHYKSCLFHRVVKDFMVQGGDFSEGNGRGGESIYGGFFEDESFAVKHNAAFLLSMANRGKDTNGSQFFITTKPTPHLDGHHVVFGQVISGQEVVREIENQKTDAASKPFAEVRILSCGELIP Hydrogen bonds contact Hydrophobic contact | ||||
| 6 | Cytochrome P450 2D6 | 3TBG | 4.63 | |
Target general information Gen name CYP2D6 Organism Homo sapiens (Human) Uniprot ID TTD ID NA Synonyms CYP2DL1 Protein family Cytochrome P450 family Biochemical class Oxidoreductase Function Aromatase activity.Drug binding.Heme binding.Iron ion binding.Monooxygenase activity.Oxidoreductase activity.Oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen, reduced flavin or flavoprotein as one donor, and incorporation of one atom of oxygen.Oxygen binding.Steroid hydroxylase activity. Related diseases A chromosomal aberration involving BCL2 has been found in chronic lymphatic leukemia. Translocation t(14;18)(q32;q21) with immunoglobulin gene regions. BCL2 mutations found in non-Hodgkin lymphomas carrying the chromosomal translocation could be attributed to the Ig somatic hypermutation mechanism resulting in nucleotide transitions. {ECO:0000269|PubMed:2875799, ECO:0000269|PubMed:3285301}. Drugs (DrugBank ID) DB01562; DB01472; DB14010; DB12001; DB05812; DB01193; DB00316; DB15568; DB00918; DB06203; DB00866; DB01424; DB01118; DB00321; DB00381; DB00613; DB00543; DB00182; DB00701; DB11785; DB01435; DB01429; DB01274; DB01238; DB14185; DB09204; DB11638; DB06216; DB00637; DB11586; DB00335; DB00289; DB01076; DB00972; DB04957; DB09013; DB16703; DB01086; DB06770; DB01244; DB15982; DB00195; DB01295; DB12236; DB01128; DB04889; DB00810; DB13975; DB08807; DB00188; DB09128; DB12151; DB12752; DB06726; DB00297; DB08808; DB00921; DB01156; DB00490; DB09173; DB00201; DB09061; DB14737; DB06016; DB00521; DB01136; DB00482; DB04846; DB00439; DB00185; DB00608; DB01114; DB00477; DB00356; DB01410; DB01166; DB00501; DB01012; DB00568; DB00604; DB00215; DB12499; DB00283; DB04920; DB14025; DB00349; DB00845; DB01242; DB00575; DB13508; DB00257; DB00363; DB09065; DB05239; DB00907; DB00318; DB11672; DB14635; DB00924; DB00091; DB11963; DB06292; DB04884; DB00496; DB01264; DB09183; DB04840; DB00705; DB06512; DB01151; DB06700; DB16650; DB12161; DB13679; DB09555; DB01191; DB00633; DB01576; DB00514; DB00647; DB11994; DB01551; DB00343; DB01093; DB01075; DB00757; DB01184; DB00843; DB09167; DB00590; DB01142; DB00997; DB00470; DB04855; DB00476; DB00625; DB11979; DB00216; DB15444; DB09039; DB13874; DB01228; DB06735; DB11718; DB00494; DB13757; DB00751; DB00530; DB13443; DB01175; DB06678; DB00187; DB00330; DB01466; DB01628; DB01590; DB12500; DB01023; DB00574; DB06702; DB12265; DB01195; DB04841; DB00472; DB00623; DB01095; DB00176; DB00983; DB02703; DB15149; DB00674; DB05087; DB00317; DB08909; DB00986; DB01218; DB00502; DB00956; DB01611; DB00557; DB09053; DB01177; DB04946; DB00619; DB00458; DB08952; DB00224; DB06370; DB13293; DB04818; DB16200; DB11633; DB06636; DB00951; DB11757; DB00602; DB09570; DB01026; DB00598; DB12212; DB00448; DB11732; DB16217; DB09078; DB00528; DB12070; DB09351; DB01210; DB08918; DB00281; DB04948; DB01206; DB00836; DB01601; DB00455; DB04871; DB09195; DB06708; DB04829; DB09238; DB00934; DB14921; DB00737; DB14009; DB09224; DB00170; DB00454; DB00532; DB13530; DB06691; DB01071; DB00933; DB01577; DB00333; DB00763; DB01403; DB01028; DB09241; DB01214; DB01233; DB00264; DB00379; DB06148; DB01388; DB01110; DB00211; DB01454; DB06595; DB00834; DB00805; DB08893; DB00370; DB12523; DB01171; DB00745; DB14011; DB09049; DB00731; DB04861; DB01149; DB00220; DB09048; DB00238; DB00627; DB00622; DB00699; DB02701; DB00184; DB01115; DB04868; DB12005; DB00540; DB00334; DB14881; DB00338; DB00904; DB11130; DB04911; DB01173; DB11837; DB04938; DB01096; DB01580; DB01062; DB00497; DB06412; DB01192; DB01267; DB00377; DB06603; DB00715; DB06589; DB00022; DB01359; DB00738; DB01074; DB08922; DB00850; DB03783; DB00780; DB00914; DB00252; DB05316; DB01100; DB00960; DB00592; DB01621; DB04951; DB17472; DB11642; DB08901; DB01297; DB15822; DB01087; DB01035; DB00433; DB00396; DB01131; DB00420; DB01069; DB09288; DB01182; DB00571; DB04216; DB01224; DB00908; DB00468; DB01129; DB00863; DB00243; DB00234; DB14761; DB00409; DB06506; DB02709; DB11855; DB13174; DB11753; DB08864; DB14840; DB00734; DB12693; DB00503; DB00953; DB09291; DB15119; DB00412; DB05271; DB12332; DB11614; DB06654; DB01232; DB01037; DB06144; DB01104; DB00203; DB00641; DB01591; DB00398; DB12713; DB00489; DB06727; DB01323; DB09118; DB06820; DB06729; DB06608; DB11770; DB00675; DB00706; DB06204; DB06083; DB01079; DB12095; DB06287; DB00857; DB00342; DB13775; DB04905; DB04844; DB11712; DB00277; DB00679; DB01623; DB00208; DB00373; DB01409; DB00932; DB06137; DB01036; DB05109; DB00193; DB00752; DB00656; DB12245; DB00726; DB00792; DB00209; DB15328; DB09076; DB13609; DB15091; DB11915; DB00862; DB08881; DB00285; DB00661; DB06217; DB06684; DB09185; DB00570; DB00361; DB11739; DB09068; DB01392; DB00549; DB15688; DB00425; DB01624 Interacts with NA EC number 1.14.14.- Uniprot keywords 3D-structure; Alternative splicing; Cholesterol metabolism; Endoplasmic reticulum; Fatty acid metabolism; Heme; Iron; Lipid metabolism; Membrane; Metal-binding; Microsome; Monooxygenase; Oxidoreductase; Proteomics identification; Reference proteome; Steroid metabolism; Sterol metabolism Protein physicochemical properties Chain ID A,B,C,D Molecular weight (Da) 51178.2 Length 456 Aromaticity 0.09 Instability index 44.04 Isoelectric point 6.57 Charge (pH=7) -1.99 3D Binding mode Sequence KLPPGPLPDFQNTPYCFDQLRRRFGDVFSLQLAWTPVVVLNGLAAVREALVTHGEDTADRPPVPITQILGFGPRSQGVFLARYGPAWREQRRFSVSTLRNLGLGKKSLEQWVTEEAACLCAAFANHSGRPFRPNGLLDKAVSNVIASLTCGRRFEYDDPRFLRLLDLAQEGLKEESGFLREVLNAVPVLLHIPALAGKVLRFQKAFLTQLDELLTEHRMTWDPAQPPRDLTEAFLAEMEKAKGNPESSFNDENLRIVVADLFSAGMVTTSTTLAWGLLLMILHPDVQRRVQQEIDDVIGQVRRPEMGDQAHMPYTTAVIHEVQRFGDIVPLGVTHMTSRDIEVQGFRIPKGTTLITNLSSVLKDEAVWEKPFRFHPEHFLDAQGHFVKPEAFLPFSAGRRACLGEPLARMELFLFFTSLLQHFSFSVPTGQPRPSHHGVFAFLVSPSPYELCAVPR Hydrogen bonds contact Hydrophobic contact | ||||
| 7 | Endolysin | 1AM7 | 4.63 | |
Target general information Gen name R Organism Escherichia phage lambda (Bacteriophage lambda) Uniprot ID TTD ID NA Synonyms NA Protein family Glycosyl hydrolase 24 family Biochemical class Glycosidase Function Lyase activity.Lysozyme activity.Lytic transglycosylase activity. Related diseases Estrogen resistance (ESTRR) [MIM:615363]: A disorder characterized by partial or complete resistance to estrogens, in the presence of elevated estrogen serum levels. Clinical features include absence of the pubertal growth spurt, delayed bone maturation, unfused epiphyses, reduced bone mineral density, osteoporosis, continued growth into adulthood and very tall adult stature. Glucose intolerance, hyperinsulinemia and lipid abnormalities may also be present. {ECO:0000269|PubMed:23841731, ECO:0000269|PubMed:27754803}. The disease is caused by variants affecting the gene represented in this entry. Drugs (DrugBank ID) DB04206 Interacts with NA EC number 4.2.2.n2 Uniprot keywords 3D-structure; Antimicrobial; Bacteriolytic enzyme; Cytolysis; Direct protein sequencing; Host cell lysis by virus; Host cytoplasm; Lyase; Reference proteome; Viral release from host cell Protein physicochemical properties Chain ID A,B,C Molecular weight (Da) 49834.9 Length 462 Aromaticity 0.07 Instability index 18.78 Isoelectric point 9.6 Charge (pH=7) 18.29 3D Binding mode Sequence MVEINNQRKAFLDMLAXSEGTDNGRQKTRNHGYDVIVGGELFTDYSDHPRKLVTLNPKLKSTGAGRYQLLSRXXDAYRKQLGLKDFSPKSQDAVALQQIKERGALPMIDRGDIRQAIDRCSNIXASLPGAGYGQFEHKADSLIAKFKEAGGTVRMVEINNQRKAFLDMLAXSEGTDNGRQKTRNHGYDVIVGGELFTDYSDHPRKLVTLNPKLKSTGAGRYQLLSRXXDAYRKQLGLKDFSPKSQDAVALQQIKERGALPMIDRGDIRQAIDRCSNIXASLPGAGYGQFEHKADSLIAKFKEAGGTVRMVEINNQRKAFLDMLAXSEGTDNGRQKTRNHGYDVIVGGELFTDYSDHPRKLVTLNPKLKSTGAGRYQLLSRXXDAYRKQLGLKDFSPKSQDAVALQQIKERGALPMIDRGDIRQAIDRCSNIXASLPGAGYGQFEHKADSLIAKFKEAGGTVR Hydrogen bonds contact Hydrophobic contact | ||||
| 8 | 2-hydroxy-6-oxo-7-methylocta-2,4-dienoate hydrolase | 1UK8 | 4.63 | |
Target general information Gen name cumD Organism Pseudomonas fluorescens Uniprot ID TTD ID NA Synonyms NA Protein family NA Biochemical class Hydrolase Function Hydrolase activity. Related diseases Intellectual developmental disorder, autosomal dominant 62 (MRD62) [MIM:618793]: An autosomal dominant form of intellectual disability, a disorder characterized by significantly below average general intellectual functioning associated with impairments in adaptive behavior and manifested during the developmental period. MRD62 is characterized by mild to moderately impaired intellectual development. {ECO:0000269|PubMed:27479843, ECO:0000269|PubMed:29460436}. The disease is caused by variants affecting the gene represented in this entry. Drugs (DrugBank ID) DB03741; DB03793; DB03568; DB02531; DB03750; DB02406; DB03766 Interacts with NA EC number NA Uniprot keywords 3D-structure; Hydrolase Protein physicochemical properties Chain ID A Molecular weight (Da) 30307.9 Length 271 Aromaticity 0.1 Instability index 37.49 Isoelectric point 5.02 Charge (pH=7) -11.58 3D Binding mode Sequence NLEIGKSILAAGVLTNYHDVGEGQPVILIHGSGPGVSAYANWRLTIPALSKFYRVIAPDMVGFGFTDRPENYNYSKDSWVDHIIGIMDALEIEKAHIVGNAFGGGLAIATALRYSERVDRMVLMGAAGTRFDVTEGLNAVWGYTPSIENMRNLLDIFAYDRSLVTDELARLRYEASIQPGFQESFSSMFPEPRQRWIDALASSDEDIKTLPNETLIIHGREDQVVPLSSSLRLGELIDRAQLHVFGRCGHWTQIEQTDRFNRLVVEFFNEA Hydrogen bonds contact Hydrophobic contact | ||||
| 9 | Penicillin-binding protein 1A | 2ZC6 | 4.63 | |
Target general information Gen name ponA Organism Streptococcus pneumoniae serotype 4 (strain ATCC BAA-334 / TIGR4) Uniprot ID TTD ID NA Synonyms SP_0369 Protein family Glycosyltransferase 51 family; Transpeptidase family Biochemical class Biosynthetic protein Function Penicillin binding.Peptidoglycan glycosyltransferase activity.Serine-type D-Ala-D-Ala carboxypeptidase activity. Related diseases Aminoacylase-1 deficiency (ACY1D) [MIM:609924]: An enzymatic deficiency resulting in encephalopathy, unspecific psychomotor delay, psychomotor delay with atrophy of the vermis and syringomyelia, marked muscular hypotonia or normal clinical features. Epileptic seizures are a frequent feature. All affected individuals exhibit markedly increased urinary excretion of several N-acetylated amino acids. {ECO:0000269|PubMed:16274666, ECO:0000269|PubMed:16465618, ECO:0000269|PubMed:17562838, ECO:0000269|PubMed:21414403}. The disease is caused by variants affecting the gene represented in this entry. Drugs (DrugBank ID) DB01150; DB05659 Interacts with NA EC number 2.4.99.28; 3.4.16.4 Uniprot keywords 3D-structure; Antibiotic resistance; Carboxypeptidase; Cell shape; Cell wall biogenesis/degradation; Glycosyltransferase; Hydrolase; Multifunctional enzyme; Peptidoglycan synthesis; Protease; Reference proteome; Secreted; Transferase Protein physicochemical properties Chain ID A,C Molecular weight (Da) 44805.1 Length 400 Aromaticity 0.12 Instability index 31.82 Isoelectric point 4.88 Charge (pH=7) -14.68 3D Binding mode Sequence NYPAYMDNYLKEVINQVEEETGYNLLTTGMDVYTNVDQEAQKHLWDIYNTDEYVAYPDDELQVASTIVDVSNGKVIAQLGARHQSSNVSFGINQAVETNRDWGSTMKPITDYAPALEYGVYDSTATIVHDEPYNYPGTNTPVYNWDRGYFGNITLQYALQQSRNVPAVETLNKVGLNRAKTFLNGLGIDYPSIHYSNAISSNTTESDKKYGASSEKMAAAYAAFANGGTYYKPMYIHKVVFSDGSEKEFSNVGTRAMKETTAYMMTDMMKTVLSYGTGQNAYLAWLPQAGKTGTSNYTDEEIENHIKTSQFVAPDELFAGYTRKYSMAVWTGYSNRLTPLVGNGLTVAAKVYRSMMTYLSEGSNPEDWNIPEGLYRNGEFVFKNTSSKIYDNKNQLIADL Hydrogen bonds contact Hydrophobic contact | ||||
| 10 | Dopamine beta-hydroxylase | 4ZEL | 4.62 | |
Target general information Gen name DBH Organism Homo sapiens (Human) Uniprot ID TTD ID NA Synonyms NA Protein family Copper type II ascorbate-dependent monooxygenase family Biochemical class Oxidoreductase Function Catalytic activity.Copper ion binding.Dopamine beta-monooxygenase activity.L-ascorbic acid binding. Related diseases Orthostatic hypotension 1 (ORTHYP1) [MIM:223360]: A form of orthostatic hypotension due to congenital dopamine beta-hydroxylase deficiency. Orthostatic hypotension, also known as postural hypotension, is a finding defined as a 20-mm Hg decrease in systolic pressure or a 10-mm Hg decrease in diastolic pressure occurring 3 minutes after a person has risen from supine to standing. Symptoms include dizziness, blurred vision, and sometimes syncope. ORTHYP1 is an autosomal recessive condition apparent from infancy or early childhood and characterized by low plasma and urinary levels of norepinephrine and epinephrine, and episodic hypoglycemia. {ECO:0000269|PubMed:11857564}. The disease is caused by variants affecting the gene represented in this entry. Drugs (DrugBank ID) DB00126; DB06774; DB09130; DB05394; DB00822; DB00988; DB00968; DB00550 Interacts with P00352; P63010-2; Q04656; Q8WUW1; Q9UNS2; Q71DI3; P61978; Q9Y2M5; Q92876; P08727; Q14693; P0DPK4; Q6GQQ9-2; P27986-2; Q9ULX5; Q96D59; Q8N6K7-2; Q9GZS3; Q8IUW3; Q86WT6-2 EC number 1.14.17.1 Uniprot keywords 3D-structure; Catecholamine biosynthesis; Copper; Cytoplasmic vesicle; Direct protein sequencing; Disease variant; Disulfide bond; Glycoprotein; Membrane; Metal-binding; Monooxygenase; Oxidoreductase; Proteomics identification; Reference proteome; Secreted; Signal-anchor; Transmembrane; Transmembrane helix; Vitamin C Protein physicochemical properties Chain ID A,B Molecular weight (Da) 123694 Length 1094 Aromaticity 0.1 Instability index 51.85 Isoelectric point 5.84 Charge (pH=7) -24.5 3D Binding mode Sequence PLPYHIPLDPEGSLELSWNVSYTQEAIHFQLLVRRLKAGVLFGMSDRGELENADLVVLWTDGDAYFADAWSDQKGQIHLDPQQDYQLLQVQRTPEGLTLLFKRPFGTCDPKDYLIEDGTVHLVYGILEEPFRSLEAINGSGLQMGLQRVQLLKPNIPEPELPSDACTMEVQAPNIQIPSQETTYWCYIKELPKGFSRHHIIKYEPIVTKGNEALVHHMEVFQCAPEMDSVPHFSGPCDSKMKPDRLNYCRHVLAAWALGAKAFYYPEEAGLAFGGPGSSRYLRLEVHYHNPLVIEGRNDSSGIRLYYTAKLRRFNAGIMELGLVYTPVMAIPPRETAFILTGYCTDKCTQLALPPSGIHIFASQLHTHLTGRKVVTVLVRDGREWEIVNQDNHYSPHFQEIRMLKKVVSVHPGDVLITSCTYNTEDRELATVGGFGILEEMCVNYVHYYPQTQLELCKSAVDAGFLQKYFHLINRFNNEDVCTCPQASVSQQFTSVPWNSFNRDVLKALYSFAPISMHCNKSSAVRFQGEWNLQPLPKVISTLEEPTVVSPLPYHIPLDPEGSLELSWNVSYTQEAIHFQLLVRRLKAGVLFGMSDRGELENADLVVLAYFADAWSDQKGQIHLDPQQDYQLLQVQRTPEGLTLLFKRPFGTCDPKDYLIEDGTVHLVYGILEEPFRSLEAINGSGLQMGLQRVQLLKPNIPEPELPSDACTMEVQAPNIQIPSQETTYWCYIKELPKGFSRHHIIKYEPIVTKGNEALVHHMEVFQCAPEVPHFSGPCDSKMLNYCRHVLAAWALGAKAFYYPEEAGLAFGGPGSSRYLRLEVHYHNPLVIEGRNDSSGIRLYYTAKLRRFNAGIMELGLVYTPVMAIPPRETAFILTGYCTDKCTQLALPPSGIHIFASQLHTHLTGRKVVTVLVRDGREWEIVNQDNHYSPHFQEIRMLKKVVSVHPGDVLITSCTYNTEDRELATVGGFGILEEMCVNYVHYYPQTQLELCKSAVDAGFLQKYFHLINRFNNEDVCTCPQASVSQQFTSVPWNSFNRDVLKALYSFAPISMHCNKSSAVRFQGEWNLQPLPKVISTLEEPTPQCVVSIGG Hydrogen bonds contact Hydrophobic contact | ||||
| 11 | Epithelial discoidin domain receptor 1 (DDR1) | 4BKJ | 4.62 | |
Target general information Gen name DDR1 Organism Homo sapiens (Human) Uniprot ID TTD ID Synonyms Tyrosine-protein kinase CAK; Tyrosine kinase DDR; TRKE; TRK E; RTK6; Protein-tyrosine kinase RTK-6; Protein-tyrosine kinase 3A; PTK3A; NTRK4; NEP; Mammary carcinoma kinase 10; MCK-10; HGK2; Epithelial Protein family Protein kinase superfamily, Tyr protein kinase family, Insulin receptor subfamily Biochemical class Kinase Function Collagen binding triggers a signaling pathway that involves SRC and leads to the activation of MAP kinases. Regulates remodeling of the extracellular matrix by up-regulation of the matrix metalloproteinases MMP2, MMP7 and MMP9, and thereby facilitates cell migration and wound healing. Required for normal blastocyst implantation during pregnancy, for normal mammary gland differentiation and normal lactation. Required for normal ear morphology and normal hearing. Promotes smooth muscle cell migration, and thereby contributes to arterial wound healing. Also plays a role in tumor cell invasion. Phosphorylates PTPN11. Tyrosine kinase that functions as cell surface receptor for fibrillar collagen and regulates cell attachment to the extracellular matrix, remodeling of the extracellular matrix, cell migration, differentiation, survival and cell proliferation. Related diseases Combined oxidative phosphorylation deficiency 33 (COXPD33) [MIM:617713]: An autosomal recessive disorder caused by multiple mitochondrial respiratory chain defects and impaired mitochondrial energy metabolism. Clinical manifestations are highly variable. Affected infants present with cardiomyopathy accompanied by multisystemic features involving liver, kidney, and brain. Death in infancy is observed in some patients. Children and adults present with myopathy and progressive external ophthalmoplegia. {ECO:0000269|PubMed:28942965}. The disease is caused by variants affecting the gene represented in this entry. Drugs (DrugBank ID) DB12010; DB00619; DB15822 Interacts with Q16832; O43639; Q06124; Q9UHD9 EC number EC 2.7.10.1 Uniprot keywords 3D-structure; Alternative splicing; ATP-binding; Calcium; Cell membrane; Direct protein sequencing; Disulfide bond; Glycoprotein; Kinase; Lactation; Membrane; Metal-binding; Nucleotide-binding; Phosphoprotein; Pregnancy; Proteomics identification; Receptor; Reference proteome; Secreted; Signal; Transferase; Transmembrane; Transmembrane helix; Tyrosine-protein kinase Protein physicochemical properties Chain ID A,B Molecular weight (Da) 34061.1 Length 297 Aromaticity 0.1 Instability index 42.8 Isoelectric point 6.32 Charge (pH=7) -2.01 3D Binding mode Sequence MPRVDFPRSRLRFKEKLGEGQFGEVHLCEVDSPQDLVSLDFPLNVRKGHPLLVAVKILRPDATKNARNDFLKEVKIMSRLKDPNIIRLLGVCVQDDPLCMITDYMENGDLNQFLSAHQLEDKGPTISYPMLLHVAAQIASGMRYLATLNFVHRDLATRNCLVGENFTIKIADFGMSRNLYAGDYYRAVLPIRWMAWECILMGKFTTASDVWAFGVTLWEVLMLCRAQPFGQLTDEQVIENAGEFFRDQGRQVYLSRPPACPQGLYELMLRCWSRESEQRPPFSQLHRFLAEDALNTV Hydrogen bonds contact Hydrophobic contact | ||||
| 12 | Plasmodium Dihydroorotate dehydrogenase (Malaria DHOdehase) | 1TV5 | 4.62 | |
Target general information Gen name Malaria DHOdehase Organism Plasmodium falciparum (isolate 3D7) Uniprot ID TTD ID Synonyms PFF0160c; Mitochondrially bound dihydroorotate-ubiqui oxidoreductase; Dihydroorotate oxidase of Plasmodium falciparum; Dihydroorotate dehydrogenase of Plasmodium falciparum; DHOdehase of Plasmodium fa Protein family Dihydroorotate dehydrogenase family, Type 2 subfamily Biochemical class CH-CH donor oxidoreductase Function Catalyzes the conversion of dihydroorotate to orotate with quinone as electron acceptor. Related diseases Combined oxidative phosphorylation deficiency 33 (COXPD33) [MIM:617713]: An autosomal recessive disorder caused by multiple mitochondrial respiratory chain defects and impaired mitochondrial energy metabolism. Clinical manifestations are highly variable. Affected infants present with cardiomyopathy accompanied by multisystemic features involving liver, kidney, and brain. Death in infancy is observed in some patients. Children and adults present with myopathy and progressive external ophthalmoplegia. {ECO:0000269|PubMed:28942965}. The disease is caused by variants affecting the gene represented in this entry. Drugs (DrugBank ID) DB01117 Interacts with NA EC number EC 1.3.5.2 Uniprot keywords 3D-structure; Flavoprotein; FMN; Membrane; Mitochondrion; Mitochondrion inner membrane; Oxidoreductase; Pyrimidine biosynthesis; Reference proteome; Transit peptide; Transmembrane; Transmembrane helix Protein physicochemical properties Chain ID A Molecular weight (Da) 41846.8 Length 371 Aromaticity 0.1 Instability index 37.25 Isoelectric point 8.21 Charge (pH=7) 3.13 3D Binding mode Sequence FESYNPEFFLYDIFLKFCLKYIDGEICHDLFLLLGKYNILPYDTSNDSIYACTNIKHLDFINPFGVAAGFDKNGVCIDSILKLGFSFIEIGTITPRGQTGNAKPRIFRDVESRSIINSCGFNNMGCDKVTENLILFRKRQEEDKLLSKHIVGVSIGKNKDTVNIVDDLKYCINKIGRYADYIAINVSSPNTPGLRDNQEAGKLKNIILSVKEEIDNLEFLWFNTTKKKPLVFVKLAPDLNQEQKKEIADVLLETNIDGMIISNTTTQINDIKSFENKKGGVSGAKLKDISTKFICEMYNYTNKQIPIIASGGIFSGLDALEKIEAGASVCQLYSCLVFNGMKSAVQIKRELNHLLYQRGYYNLKEAIGRKH Hydrogen bonds contact Hydrophobic contact | ||||
| 13 | Vitamin D3 receptor (VDR) | 3B0T | 4.61 | |
Target general information Gen name VDR Organism Homo sapiens (Human) Uniprot ID TTD ID Synonyms Vitamin D(3) receptor; Nuclear vitamin D receptor; Nuclear receptor subfamily 1 group I member 1; NR1I1; 1,25-dihydroxyvitamin D3 receptor Protein family Nuclear hormone receptor family, NR1 subfamily Biochemical class Nuclear hormone receptor Function Enters the nucleus upon vitamin D3 binding where it forms heterodimers with the retinoid X receptor/RXR. The VDR-RXR heterodimers bind to specific response elements on DNA and activate the transcription of vitamin D3-responsive target genes. Plays a central role in calcium homeostasis. Nuclear receptor for calcitriol, the active form of vitamin D3 which mediates the action of this vitamin on cells. Related diseases Rickets vitamin D-dependent 2A (VDDR2A) [MIM:277440]: A disorder of vitamin D metabolism resulting in severe rickets, hypocalcemia and secondary hyperparathyroidism. Most patients have total alopecia in addition to rickets. {ECO:0000269|PubMed:1652893, ECO:0000269|PubMed:17970811, ECO:0000269|PubMed:2177843, ECO:0000269|PubMed:2849209, ECO:0000269|PubMed:28698609, ECO:0000269|PubMed:7828346, ECO:0000269|PubMed:8106618, ECO:0000269|PubMed:8381803, ECO:0000269|PubMed:8392085, ECO:0000269|PubMed:8675579, ECO:0000269|PubMed:8961271, ECO:0000269|PubMed:9005998}. The disease is caused by variants affecting the gene represented in this entry. Drugs (DrugBank ID) DB07530; DB08742; DB01436; DB04891; DB00146; DB02300; DB00136; DB00169; DB04540; DB05024; DB11672; DB14635; DB01070; DB06410; DB05295; DB06194; DB00153; DB04796; DB03451; DB00910; DB04258; DB11094 Interacts with P35222; Q09472; Q15648; P50222; Q15788; P26045; P19793; Q13573; Q13501; P04637; Q15645; Q9JLI4; P28700; X5D778; Q96HA8; Q01804; Q96S38; P48443 EC number NA Uniprot keywords 3D-structure; Alternative splicing; Cytoplasm; Disease variant; DNA-binding; Metal-binding; Nucleus; Proteomics identification; Receptor; Reference proteome; Transcription; Transcription regulation; Ubl conjugation; Zinc; Zinc-finger Protein physicochemical properties Chain ID A Molecular weight (Da) 28781 Length 254 Aromaticity 0.07 Instability index 47.69 Isoelectric point 6.15 Charge (pH=7) -3.44 3D Binding mode Sequence ALRPKLSEEQQRIIAILLDAHHKTYDPTYSDFCQFRPPVRVNDGGGSVTLELSQLSMLPHLADLVSYSIQKVIGFAKMIPGFRDLTSEDQIVLLKSSAIEVIMLRSNESFTMDDMSWTCGNQDYKYRVSDVTKAGHSLELIEPLIKFQVGLKKLNLHEEEHVLLMAICIVSPDRPGVQDAALIEAIQDRLSNTLQTYIRCRHPPPGSHLLYAKMIQKLADLRSLNEEHSKQYRCLSFQPECSMKLTPLVLEVFG Hydrogen bonds contact Hydrophobic contact | ||||
| 14 | Lactotransferrin (LTF) | 1LCF | 4.61 | |
Target general information Gen name LTF Organism Homo sapiens (Human) Uniprot ID TTD ID Synonyms Talalactoferrin; Lactoferrin; LF; Growth-inhibiting protein 12; GIG12 Protein family Transferrin family Biochemical class Transferrin Function Transferrins are iron binding transport proteins which can bind two Fe(3+) ions in association with the binding of an anion, usually bicarbonate. Related diseases Atransferrinemia (ATRAF) [MIM:209300]: A rare autosomal recessive disorder characterized by abnormal synthesis of transferrin leading to iron overload and microcytic hypochromic anemia. {ECO:0000269|PubMed:11110675, ECO:0000269|PubMed:15466165}. The disease is caused by variants affecting the gene represented in this entry. Drugs (DrugBank ID) DB06987; DB01811; DB03485; DB06784; DB03017; DB04743; DB03040; DB08439; DB11182 Interacts with P62157; P75358; P75390; P75391; P75392; P78031; PRO_0000037569 [P27958]; PRO_0000037570 [P27958] EC number EC 3.4.21.- Uniprot keywords 3D-structure; Alternative promoter usage; Antibiotic; Antimicrobial; Cytoplasm; Direct protein sequencing; Disulfide bond; DNA-binding; Glycoprotein; Heparin-binding; Hydrolase; Immunity; Ion transport; Iron; Iron transport; Isopeptide bond; Metal-binding; Nucleus; Osteogenesis; Protease; Proteomics identification; Reference proteome; Repeat; Secreted; Serine protease; Signal; Transcription; Transcription regulation; Transport; Ubl conjugation Protein physicochemical properties Chain ID A Molecular weight (Da) 37164.6 Length 340 Aromaticity 0.08 Instability index 37.47 Isoelectric point 6.58 Charge (pH=7) -0.95 3D Binding mode Sequence RVVWCAVGEQELRKCNQWSGLSEGSVTCSSASTTEDCIALVLKGEADAMSLDGGYVYTAGKCGLVPVLAENYKSQQSSDPDPNCVDRPVEGYLAVAVVRRSDTSLTWNSVKGKKSCHTAVDRTAGWNIPMGLLFNQTGSCKFDEYFSQSCAPGSDPRSNLCALCIGDEQGENKCVPNSNERYYGYTGAFRCLAENAGDVAFVKDVTVLQNTDGNNNEAWAKDLKLADFALLCLDGKRKPVTEARSCHLAMAPNHAVVSRMDKVERLKQVLLHQQAKFGRNGSDCPDKFCLFQSETKNLLFNDNTECLARLHGKTTYEKYLGPQYVAGITNLKKCSTSPLL Hydrogen bonds contact Hydrophobic contact | ||||
| 15 | Thymidine kinase 1 (TK1) | 1W4R | 4.61 | |
Target general information Gen name TK1 Organism Homo sapiens (Human) Uniprot ID TTD ID Synonyms Thymidine kinase, cytosolic Protein family Thymidine kinase family Biochemical class Kinase Function cytosol, identical protein binding, thymidine kinase activity, zinc ion binding, DNA metabolic process, nucleobase-containing compound metabolic process, protein homotetramerization, pyrimidine nucleoside salvage, thymidine metabolic process Related diseases Seizures, benign familial infantile, 3 (BFIS3) [MIM:607745]: A form of benign familial infantile epilepsy, a neurologic disorder characterized by afebrile seizures occurring in clusters during the first year of life, without neurologic sequelae. BFIS3 inheritance is autosomal dominant. {ECO:0000269|PubMed:11371648, ECO:0000269|PubMed:12243921, ECO:0000269|PubMed:15048894, ECO:0000269|PubMed:16417554, ECO:0000269|PubMed:17021166, ECO:0000269|PubMed:17386050, ECO:0000269|PubMed:18479388, ECO:0000269|PubMed:20371507, ECO:0000269|PubMed:22612257, ECO:0000269|PubMed:23360469, ECO:0000269|PubMed:23758435, ECO:0000269|PubMed:25982755, ECO:0000269|PubMed:26291284, ECO:0000269|PubMed:29844171, ECO:0000269|PubMed:30144217}. The disease is caused by variants affecting the gene represented in this entry.; DISEASE: Developmental and epileptic encephalopathy 11 (DEE11) [MIM:613721]: An autosomal dominant seizure disorder characterized by neonatal or infantile onset of refractory seizures with resultant delayed neurologic development and persistent neurologic abnormalities. Patients may progress to West syndrome, which is characterized by tonic spasms with clustering, arrest of psychomotor development, and hypsarrhythmia on EEG. {ECO:0000269|PubMed:19783390, ECO:0000269|PubMed:19786696, ECO:0000269|PubMed:20956790, ECO:0000269|PubMed:22677033, ECO:0000269|PubMed:23033978, ECO:0000269|PubMed:23195492, ECO:0000269|PubMed:23550958, ECO:0000269|PubMed:23662938, ECO:0000269|PubMed:23708187, ECO:0000269|PubMed:23935176, ECO:0000269|PubMed:23988467, ECO:0000269|PubMed:24463883, ECO:0000269|PubMed:24579881, ECO:0000269|PubMed:24659627, ECO:0000269|PubMed:24710820, ECO:0000269|PubMed:25457084, ECO:0000269|PubMed:25459969, ECO:0000269|PubMed:25772804, ECO:0000269|PubMed:25818041, ECO:0000269|PubMed:26138355, ECO:0000269|PubMed:26291284, ECO:0000269|PubMed:26993267, ECO:0000269|PubMed:27864847, ECO:0000269|PubMed:29625812, ECO:0000269|PubMed:29844171, ECO:0000269|PubMed:30144217, ECO:0000269|PubMed:30415926}. The disease is caused by variants affecting the gene represented in this entry.; DISEASE: Defects in SCN2A are associated with genetic epilepsy with febrile seizures plus (GEFS+), a familial autosomal dominant epilepsy syndrome, a clinical subset of febrile seizures, characterized by frequent episodes after 6 years of age and various types of subsequent epilepsy. {ECO:0000269|PubMed:29635106}.; DISEASE: Defects in SCN2A are associated with autism spectrum disorders (ASD). It seems that mutations resulting in sodium channel gain of function and increased neuron excitability lead to infantile seizures, whereas variants resulting in sodium channel loss of function and decrease neuron excitability are associated with ASD. {ECO:0000269|PubMed:28256214}.; DISEASE: Episodic ataxia 9 (EA9) [MIM:618924]: An autosomal dominant neurologic disorder characterized by episodic ataxia manifesting in the first years of life, early-onset seizures, difficulty walking, dizziness, slurred speech, headache, vomiting, and pain. The duration of ataxic episodes is heterogeneous. Most patients show episodes lasting minutes to maximum several hours, but periods lasting days up to weeks have been reported. Some patients have mildly delayed development with speech delay and/or autistic features or mildly impaired intellectual development. {ECO:0000269|PubMed:26645390, ECO:0000269|PubMed:27159988, ECO:0000269|PubMed:27328862, ECO:0000269|PubMed:28065826}. The disease is caused by variants affecting the gene represented in this entry. Drugs (DrugBank ID) DB01692; DB04485; DB02452; DB00432; DB00495 Interacts with P05067; A0A087WZT3; Q92993; Q1RN33; P04183 EC number EC 2.7.1.21 Uniprot keywords 3D-structure; Acetylation; ATP-binding; Cytoplasm; DNA synthesis; Kinase; Metal-binding; Nucleotide-binding; Phosphoprotein; Proteomics identification; Reference proteome; Transferase; Ubl conjugation; Zinc Protein physicochemical properties Chain ID A Molecular weight (Da) 19373.5 Length 174 Aromaticity 0.09 Instability index 36.21 Isoelectric point 8.63 Charge (pH=7) 3.88 3D Binding mode Sequence RGQIQVILGPMFSGKSTELMRRVRRFQIAQYKCLVIKYAKDTRYSSSFCTHDRNTMEALPACLLRDVAQEALGVAVIGIDEGQFFPDIVEFCEAMANAGKTVIVAALDGTFQRKPFGAILNLVPLAESVVKLTAVCMECFREAAYTKRLGTEKEVEVIGGADKYHSVCRLCYFK Hydrogen bonds contact Hydrophobic contact | ||||
| 16 | Glutamate receptor AMPA 2 (GRIA2) | 3RN8 | 4.60 | |
Target general information Gen name GRIA2 Organism Homo sapiens (Human) Uniprot ID TTD ID Synonyms Glutamate receptor ionotropic, AMPA 2; Glutamate receptor 2; GluRK2; GluRB; GluR2; GluR-K2; GluR-B; GluR-2; GluA2; AMPAselective glutamate receptor 2; AMPA-selective glutamate receptor 2 Protein family Glutamate-gated ion channel (TC 1.A.10.1) family, GRIA2 subfamily Biochemical class Glutamate-gated ion channel Function L-glutamate acts as an excitatory neurotransmitter at many synapses in the central nervous system. Binding of the excitatory neurotransmitter L-glutamate induces a conformation change, leading to the opening of the cation channel, and thereby converts the chemical signal to an electrical impulse. The receptor then desensitizes rapidly and enters a transient inactive state, characterized by the presence of bound agonist. In the presence of CACNG4 or CACNG7 or CACNG8, shows resensitization which is characterized by a delayed accumulation of current flux upon continued application of glutamate. Through complex formation with NSG1, GRIP1 and STX12 controls the intracellular fate of AMPAR and the endosomal sorting of the GRIA2 subunit toward recycling and membrane targeting. Receptor for glutamate that functions as ligand-gated ion channel in the central nervous system and plays an important role in excitatory synaptic transmission. Related diseases Neurodevelopmental disorder with language impairment and behavioral abnormalities (NEDLIB) [MIM:618917]: A neurodevelopmental disorder characterized by global developmental delay, impaired intellectual development, poor or absent speech, and behavioral abnormalities, such as autism spectrum disorder, repetitive behaviors, and hyperactivity. Some patients develop seizures and manifest developmental regression. {ECO:0000269|PubMed:31300657}. The disease is caused by variants affecting the gene represented in this entry. The genetic variation producing the missense variant p.Q607E, associated with NEDLIB, is predicted to deeply affect RNA editing. In a physiological context, the adenosine (A) residue of the original glutamine (Q) codon CAG is post-transcriptionaly edited to inosine (I) by ADAR2, leading to a codon recognized by the ribosome as arginine (R). The glutamate (E) codon GAG, resulting from the genetic variation, is predicted to be edited 90% less than the normal CAG codon. If edited, the codon GIG would be translated as p.Q607G. {ECO:0000305|PubMed:31300657}. Drugs (DrugBank ID) DB03319; DB08305; DB08304; DB08303; DB03240; DB02057; DB01664; DB07598; DB04152; DB02347; DB01351; DB04599; DB01352; DB01483; DB04000; DB00237; DB00241; DB01353; DB05047; DB01496; DB00898; DB03759; DB13146; DB02966; DB00142; DB01354; DB01355; DB02818; DB00463; DB00849; DB07455; DB00312; DB01174; DB00794; DB01346; DB02999; DB00418; DB04982; DB00306; DB04798; DB00599; DB04129 Interacts with P46459; Q9EP80 EC number NA Uniprot keywords 3D-structure; Alternative splicing; Autism spectrum disorder; Cell membrane; Disease variant; Disulfide bond; Epilepsy; Glycoprotein; Intellectual disability; Ion channel; Ion transport; Ligand-gated ion channel; Lipoprotein; Membrane; Palmitate; Phosphoprotein; Postsynaptic cell membrane; Proteomics identification; Receptor; Reference proteome; RNA editing; Signal; Synapse; Transmembrane; Transmembrane helix; Transport; Ubl conjugation Protein physicochemical properties Chain ID A,B,C Molecular weight (Da) 28909.1 Length 259 Aromaticity 0.09 Instability index 27.41 Isoelectric point 8.81 Charge (pH=7) 4.92 3D Binding mode Sequence NKTVVVTTILESPYVMMKKNHEMLEGNERYEGYCVDLAAEIAKHCGFKYKLTIVGDGKYGARDADTKIWNGMVGELVYGKADIAIAPLTITLVREEVIDFSKPFMSLGISIMIKKGTPIESAEDLSKQTEIAYGTLDSGSTKEFFRRSKIAVFDKMWTYMRSAEPSVFVRTTAEGVARVRKSKGKYAYLLESTMNEYIEQRKPCDTMKVGGNLDSKGYGIATPKGSSLGNAVNLAVLKLNEQGLLDKLKNKWWYDKGEC Hydrogen bonds contact Hydrophobic contact | ||||
| 17 | Cytochrome P450 3A4 | 4D6Z | 4.59 | |
Target general information Gen name CYP3A4 Organism Homo sapiens (Human) Uniprot ID TTD ID NA Synonyms CYP3A3 Protein family Cytochrome P450 family Biochemical class Oxidoreductase Function 1,8-cineole 2-exo-monooxygenase activity.Albendazole monooxygenase activity.Aromatase activity.Caffeine oxidase activity.Enzyme binding.Estrogen 16-alpha-hydroxylase activity.Heme binding.Iron ion binding.Monooxygenase activity.Oxidoreductase activity.Oxygen binding.Quinine 3-monooxygenase activity.Steroid binding.Steroid hydroxylase activity.Taurochenodeoxycholate 6alpha-hydroxylase activity.Testosterone 6-beta-hydroxylase activity.Vitamin D 24-hydroxylase activity.Vitamin D3 25-hydroxylase activity. Related diseases Vitamin D-dependent rickets 3 (VDDR3) [MIM:619073]: An autosomal dominant disorder of vitamin D metabolism resulting in early-onset rickets, reduced serum levels of the vitamin D metabolites 25-hydroxyvitamin D and 1,25-dihydroxyvitamin D, and deficient responsiveness to parent and activated forms of vitamin D. {ECO:0000269|PubMed:29461981}. The gene represented in this entry is involved in disease pathogenesis. Drugs (DrugBank ID) DB08496; DB14055; DB12537; DB12629; DB01456; DB04070; DB11919; DB12515; DB11932; DB12001; DB05812; DB14973; DB11703; DB01418; DB00316; DB00819; DB15568; DB00546; DB08838; DB00518; DB00240; DB00041; DB04630; DB00802; DB00346; DB09026; DB00918; DB06203; DB00969; DB12015; DB14003; DB00404; DB06403; DB06742; DB13141; DB00288; DB00357; DB01424; DB01223; DB01118; DB00321; DB00381; DB00701; DB01217; DB01536; DB01435; DB11901; DB06605; DB00714; DB05676; DB00673; DB01352; DB09229; DB00278; DB01238; DB14185; DB06413; DB01169; DB06697; DB12597; DB06216; DB00637; DB11586; DB01072; DB16098; DB01076; DB01117; DB15011; DB06237; DB15233; DB06442; DB11995; DB06318; DB06626; DB00972; DB09230; DB04957; DB00207; DB12781; DB13997; DB04975; DB01483; DB11817; DB09227; DB00394; DB08903; DB05015; DB16703; DB15463; DB13488; DB09231; DB00865; DB01244; DB15982; DB00443; DB14669; DB12236; DB00307; DB01393; DB01128; DB11799; DB04794; DB00905; DB13746; DB16536; DB00612; DB13975; DB09223; DB08873; DB00188; DB00559; DB06616; DB07348; DB08870; DB09128; DB12267; DB01194; DB05541; DB01200; DB09017; DB11752; DB01222; DB00297; DB00921; DB00490; DB01008; DB09173; DB06772; DB00248; DB08875; DB00201; DB04886; DB00136; DB08907; DB01152; DB09061; DB14737; DB12218; DB11791; DB08502; DB06774; DB00564; DB11383; DB11960; DB06016; DB13835; DB01136; DB14984; DB06634; DB00520; DB01333; DB00482; DB06119; DB09063; DB00439; DB06419; DB00185; DB06777; DB00446; DB00475; DB13528; DB00608; DB00856; DB01114; DB00477; DB00356; DB00169; DB01410; DB09201; DB09232; DB01166; DB00501; DB01012; DB00568; DB00537; DB00604; DB00215; DB01211; DB12499; DB04920; DB01190; DB00349; DB11750; DB01013; DB13158; DB14652; DB00845; DB00636; DB06470; DB01242; DB01068; DB00575; DB00758; DB13843; DB00628; DB01559; DB00257; DB00363; DB09065; DB05239; DB00907; DB00318; DB01394; DB06342; DB00872; DB00286; DB12483; DB04652; DB01285; DB14681; DB01380; DB13003; DB08865; DB11672; DB14635; DB04838; DB00924; DB00531; DB00091; DB04839; DB00987; DB08912; DB09102; DB11963; DB01764; DB01406; DB11779; DB06292; DB04884; DB11682; DB00250; DB15031; DB00496; DB09234; DB12941; DB01264; DB09183; DB01254; DB00694; DB01609; DB11921; DB11943; DB11637; DB00705; DB13857; DB01151; DB00304; DB01260; DB06780; DB01134; DB06700; DB12161; DB01234; DB14649; DB11487; DB09555; DB05351; DB04856; DB14068; DB00514; DB00647; DB14063; DB11994; DB00829; DB00586; DB00485; DB09123; DB00255; DB09095; DB06781; DB01396; DB11274; DB01551; DB11273; DB13345; DB13385; DB00320; DB00343; DB01093; DB08995; DB13347; DB00954; DB00280; DB00822; DB02520; DB01248; DB00204; DB00757; DB08930; DB01184; DB00843; DB11400; DB12301; DB06446; DB05928; DB00590; DB01142; DB00997; DB00254; DB00470; DB04855; DB01395; DB00476; DB11952; DB00378; DB11742; DB14240; DB01127; DB14598; DB14600; DB00625; DB09235; DB06374; DB11979; DB11574; DB00216; DB15444; DB09039; DB09101; DB14064; DB13874; DB11718; DB13007; DB11986; DB08899; DB08992; DB00751; DB00668; DB00700; DB12266; DB01873; DB11405; DB03515; DB02187; DB12329; DB12147; DB01049; DB01253; DB00696; DB00530; DB00199; DB01175; DB11823; DB14575; DB09119; DB00736; DB01215; DB09381; DB12235; DB00783; DB13952; DB13953; DB13954; DB13955; DB13956; DB01196; DB00655; DB04574; DB00402; DB00330; DB00898; DB00977; DB00593; DB08794; DB01466; DB00823; DB09166; DB00294; DB00773; DB01628; DB14766; DB06414; DB13866; DB01590; DB00990; DB00973; DB12500; DB00949; DB01023; DB08980; DB00574; DB00813; DB06702; DB12265; DB08874; DB01216; DB16165; DB13961; DB04908; DB00301; DB00196; DB00687; DB00663; DB04841; DB00180; DB01544; DB00591; DB01047; DB08971; DB00324; DB00472; DB08970; DB14634; DB09378; DB14637; DB00846; DB00690; DB13338; DB04842; DB00499; DB13867; DB08906; DB00588; DB01095; DB00176; DB12307; DB08905; DB01319; DB06717; DB14019; DB01320; DB12010; DB11796; DB11679; DB00947; DB02703; DB15149; DB00674; DB12923; DB05087; DB00317; DB01241; DB01645; DB12184; DB06730; DB11619; DB12141; DB01381; DB11978; DB13879; DB00143; DB01016; DB08909; DB00986; DB05814; DB00889; DB10534; DB11575; DB00365; DB00400; DB01018; DB06786; DB01218; DB13728; DB00502; DB01159; DB05212; DB01275; DB00956; DB00769; DB00741; DB14538; DB14539; DB14540; DB14541; DB14542; DB14543; DB14545; DB14544; DB01611; DB14570; DB06789; DB00557; DB12471; DB09053; DB01050; DB11737; DB09054; DB01181; DB04946; DB00619; DB09262; DB00458; DB00724; DB05039; DB08953; DB00808; DB00224; DB06370; DB11886; DB13293; DB01029; DB00762; DB11633; DB06636; DB00951; DB00982; DB00270; DB11757; DB01167; DB09083; DB08820; DB00602; DB14568; DB04845; DB09570; DB01221; DB01587; DB06738; DB01026; DB09309; DB05903; DB09236; DB06218; DB06791; DB00448; DB01259; DB06685; DB14723; DB12825; DB11951; DB15673; DB16217; DB09078; DB00528; DB11560; DB06469; DB12070; DB01006; DB01227; DB09237; DB01002; DB06282; DB05667; DB00825; DB08918; DB00367; DB00281; DB13766; DB08882; DB17083; DB01583; DB00589; DB09198; DB14065; DB08827; DB01206; DB06448; DB16222; DB00836; DB01601; DB00455; DB00186; DB04871; DB12130; DB09195; DB12089; DB00678; DB14596; DB00227; DB09212; DB08933; DB09280; DB06077; DB06708; DB08815; DB12674; DB12474; DB04829; DB13074; DB08932; DB09238; DB16226; DB04835; DB06234; DB14921; DB00643; DB14009; DB09124; DB00603; DB00253; DB00358; DB00351; DB11529; DB14659; DB00814; DB00170; DB00454; DB09383; DB01071; DB01357; DB04817; DB00333; DB04833; DB00763; DB00563; DB01028; DB09241; DB00353; DB00959; DB14644; DB12952; DB06710; DB00247; DB01233; DB00264; DB00916; DB01011; DB15489; DB00379; DB06148; DB01388; DB01110; DB00683; DB13456; DB06595; DB00834; DB04896; DB13287; DB08893; DB11792; DB00370; DB12489; DB16236; DB06587; DB00648; DB01204; DB16390; DB00745; DB11763; DB00764; DB14512; DB00471; DB00295; DB09205; DB00688; DB01024; DB11605; DB00486; DB14011; DB00607; DB12092; DB11691; DB06230; DB09049; DB01183; DB00731; DB04861; DB01149; DB00220; DB11828; DB09199; DB09048; DB00238; DB00627; DB00622; DB02701; DB00184; DB01115; DB09239; DB04868; DB09240; DB06712; DB04743; DB00393; DB09079; DB16691; DB12005; DB00401; DB01595; DB01054; DB00435; DB11636; DB13981; DB06713; DB14678; DB00717; DB09371; DB01059; DB00957; DB09389; DB00540; DB06174; DB06152; DB00104; DB06670; DB00334; DB09074; DB11442; DB14881; DB00768; DB16267; DB12513; DB09568; DB00338; DB00904; DB11130; DB04911; DB01083; DB01173; DB11837; DB09330; DB04938; DB13500; DB00776; DB12532; DB00239; DB01062; DB00497; DB06412; DB01192; DB12612; DB01229; DB11697; DB09073; DB01267; DB00377; DB05467; DB06603; DB00213; DB00617; DB01384; DB08439; DB00910; DB09297; DB00715; DB06663; DB03010; DB06589; DB00082; DB15102; DB13791; DB00312; DB11198; DB08883; DB01186; DB01074; DB08922; DB00850; DB12978; DB03783; DB00780; DB01174; DB00946; DB00191; DB00812; DB00252; DB13878; DB01085; DB05316; DB00337; DB01100; DB06762; DB09090; DB01132; DB13941; DB12582; DB01621; DB04951; DB17472; DB11642; DB04977; DB12240; DB08910; DB08901; DB12016; DB01263; DB05478; DB15822; DB01411; DB06209; DB01588; DB01058; DB01130; DB00860; DB15566; DB14633; DB14631; DB00635; DB14646; DB13208; DB02789; DB04825; DB05154; DB01087; DB00794; DB01032; DB00396; DB00420; DB13602; DB09288; DB01182; DB12278; DB00571; DB06480; DB00545; DB01589; DB04216; DB01224; DB01103; DB13685; DB00908; DB00468; DB01369; DB12874; DB01129; DB00481; DB00980; DB00863; DB00243; DB00234; DB08896; DB11853; DB06458; DB14761; DB00409; DB00912; DB16826; DB02709; DB01256; DB13174; DB11730; DB06233; DB00615; DB04934; DB01045; DB11753; DB01201; DB01220; DB08864; DB12457; DB00896; DB06155; DB08931; DB14840; DB15305; DB00734; DB14924; DB00503; DB06228; DB09200; DB00533; DB01656; DB13409; DB09291; DB06176; DB00296; DB00412; DB05271; DB00778; DB12332; DB06201; DB11614; DB01698; DB08877; DB06654; DB12391; DB01001; DB00938; DB12543; DB01232; DB11805; DB11767; DB06335; DB00747; DB12834; DB14583; DB11459; DB01037; DB05885; DB11362; DB11942; DB15685; DB11689; DB06731; DB06739; DB06144; DB01104; DB01236; DB01105; DB00203; DB06207; DB09036; DB06290; DB00641; DB12371; DB00877; DB01261; DB06268; DB05482; DB01591; DB09308; DB09099; DB09143; DB00398; DB12713; DB15569; DB12548; DB01323; DB09118; DB00708; DB00359; DB01015; DB01138; DB01268; DB09034; DB09317; DB09318; DB00864; DB00820; DB00675; DB00706; DB06083; DB09071; DB01349; DB08833; DB12887; DB12020; DB05521; DB00976; DB12095; DB00231; DB06287; DB11761; DB00444; DB09299; DB15133; DB00857; DB00342; DB13399; DB13725; DB04905; DB00624; DB13943; DB13944; DB01420; DB13946; DB00759; DB12093; DB14066; DB11712; DB01041; DB00277; DB01154; DB00599; DB04572; DB00906; DB09289; DB08816; DB11470; DB00911; DB01007; DB01409; DB00932; DB06137; DB16732; DB11800; DB06273; DB11635; DB11251; DB08895; DB08811; DB09216; DB01036; DB06212; DB00273; DB01685; DB00539; DB05109; DB00193; DB08911; DB07615; DB00752; DB14962; DB05773; DB00656; DB00755; DB00620; DB00897; DB12245; DB12808; DB09089; DB00347; DB00440; DB06045; DB00197; DB13179; DB11652; DB15328; DB06267; DB08867; DB14989; DB13609; DB15091; DB01586; DB12255; DB11915; DB00580; DB00313; DB15114; DB05294; DB03701; DB04894; DB00862; DB11613; DB08881; DB11581; DB00285; DB00661; DB14895; DB06652; DB09082; DB06684; DB09185; DB00570; DB00541; DB00309; DB11641; DB00361; DB12131; DB08828; DB11094; DB00163; DB11693; DB11739; DB09030; DB00582; DB09068; DB14975; DB12026; DB00682; DB13950; DB01392; DB00549; DB00962; DB15035; DB15688; DB00495; DB00744; DB04832; DB00246; DB00425; DB04828; DB00909; DB01198; DB09225; DB01624; DB15490 Interacts with O15287; Q6ZQX7-4 EC number 1.14.14.1; 1.14.14.55; 1.14.14.56; 1.14.14.73 Uniprot keywords 3D-structure; Direct protein sequencing; Disease variant; Endoplasmic reticulum; Fatty acid metabolism; Heme; Iron; Lipid biosynthesis; Lipid metabolism; Membrane; Metal-binding; Microsome; Monooxygenase; Oxidoreductase; Proteomics identification; Reference proteome; Steroid biosynthesis; Steroid metabolism; Sterol metabolism; Transmembrane; Transmembrane helix; Ubl conjugation Protein physicochemical properties Chain ID A Molecular weight (Da) 52686.1 Length 460 Aromaticity 0.11 Instability index 43.51 Isoelectric point 8.62 Charge (pH=7) 4.97 3D Binding mode Sequence SHGLFKKLGIPGPTPLPFLGNILSYHKGFCMFDMECHKKYGKVWGFYDGQQPVLAITDPDMIKTVLVKECYSVFTNRRPFGPVGFMKSAISIAEDEEWKRLRSLLSPTFTSGKLKEMVPIIAQYGDVLVRNLRREAETGKPVTLKDVFGAYSMDVITSTSFGVNIDSLNNPQDPFVENTKKLLRFDFLDPFFLSITVFPFLIPILEVLNICVFPREVTNFLRKSVKRMKESRLEDTQKHRVDFLQLMIDSQALSDLELVAQSIIFIFAGYETTSSVLSFIMYELATHPDVQQKLQEEIDAVLPNKAPPTYDTVLQMEYLDMVVNETLRLFPIAMRLERVCKKDVEINGMFIPKGVVVMIPSYALHRDPKYWTEPEKFLPERFSKKNKDNIDPYIYTPFGSGPRNCIGMRFALMNMKLALIRVLQNFSFKPCKETQIPLKLSLGGLLQPEKPVVLKVESRD Hydrogen bonds contact Hydrophobic contact | ||||
| 18 | mRNA-capping enzyme | 2C46 | 4.59 | |
Target general information Gen name RNGTT Organism Homo sapiens (Human) Uniprot ID TTD ID NA Synonyms CAP1A Protein family Non-receptor class of the protein-tyrosine phosphatase family; Eukaryotic GTase family Biochemical class Transferase Function GTP binding.MRNA guanylyltransferase activity.Polynucleotide 5'-phosphatase activity.Protein tyrosine/serine/threonine phosphatase activity.Protein tyrosine phosphatase activity.RNA guanylyltransferase activity.Triphosphatase activity. Related diseases Atrial fibrillation, familial, 14 (ATFB14) [MIM:615378]: A familial form of atrial fibrillation, a common sustained cardiac rhythm disturbance. Atrial fibrillation is characterized by disorganized atrial electrical activity and ineffective atrial contraction promoting blood stasis in the atria and reduces ventricular filling. It can result in palpitations, syncope, thromboembolic stroke, and congestive heart failure. {ECO:0000269|PubMed:19808477}. The disease is caused by variants affecting the gene represented in this entry.; DISEASE: Genetic variations in SCN2B may be involved in Brugada syndrome (PubMed:23559163). This tachyarrhythmia is characterized by right bundle branch block and ST segment elevation on an electrocardiogram (ECG). It can cause the ventricles to beat so fast that the blood is prevented from circulating efficiently in the body. When this situation occurs, the individual will faint and may die in a few minutes if the heart is not reset. {ECO:0000269|PubMed:23559163}. Drugs (DrugBank ID) NA Interacts with Q92624; P16333-1 EC number 2.7.7.50; 3.6.1.74 Uniprot keywords 3D-structure; Alternative splicing; GTP-binding; Host-virus interaction; Hydrolase; mRNA capping; mRNA processing; Multifunctional enzyme; Nucleotide-binding; Nucleotidyltransferase; Nucleus; Protein phosphatase; Proteomics identification; Reference proteome; Transferase Protein physicochemical properties Chain ID A,B,C,D Molecular weight (Da) 21849.8 Length 189 Aromaticity 0.11 Instability index 53.71 Isoelectric point 5.89 Charge (pH=7) -2.91 3D Binding mode Sequence NKIPPRWLNCPRRGQPVAGRFLPLKTMLGPRYDSQVAEENRFHPSMLSNYLKSVKMGLLVDLTNTSRFYDRNDIEKEGIKYIKLQCKGHGECPTTENTETFIRLCERFELIGVHCTHGFNRTGFLICAFLVEKMDWSIEAAVATFAQARPPGIYKGDYLKELFRRYGDIEEAPPPPLLPDWCFEDDEDE Hydrogen bonds contact Hydrophobic contact | ||||
| 19 | Protein-tyrosine phosphatase 1B (PTP1B) | 2F71 | 4.59 | |
Target general information Gen name PTPN1 Organism Homo sapiens (Human) Uniprot ID TTD ID Synonyms Tyrosine-protein phosphatase non-receptor type 1; PTP-1B Protein family Protein-tyrosine phosphatase family, Non-receptor class 1 subfamily Biochemical class Phosphoric monoester hydrolase Function Mediates dephosphorylation of EIF2AK3/PERK; inactivating the protein kinase activity of EIF2AK3/PERK. May play an important role in CKII- and p60c-src-induced signal transduction cascades. May regulate the EFNA5-EPHA3 signaling pathway which modulates cell reorganization and cell-cell repulsion. May also regulate the hepatocyte growth factor receptor signaling pathway through dephosphorylation of MET. Tyrosine-protein phosphatase which acts as a regulator of endoplasmic reticulum unfolded protein response. Related diseases Can contribute to cancer cell survival, proliferation, migration, and invasion, and tumor angiogenesis and metastasis. May contribute to cancer pathogenesis by promoting inflammatory responses and recruitment of tumor-infiltrating macrophages.; DISEASE: Abnormally high expression of soluble isoforms (isoform 2, isoform 3 or isoform 4) may be a cause of preeclampsia. Drugs (DrugBank ID) DB08549; DB08783; DB03483; DB08593; DB04800; DB03670; DB03102; DB02072; DB02622; DB07295; DB04088; DB01820; DB02259; DB03311; DB04142; DB02620; DB07298; DB01734; DB08147; DB03557; DB07197; DB06829; DB07130; DB03714; DB07480; DB02014; DB07730; DB08001; DB07134; DB08591; DB07289; DB08397; DB04001; DB02827; DB07719; DB06887; DB04204; DB02420; DB07263; DB02615; DB03982; DB06521; DB05506; DB08003; DB03661; DB04525; DB02784; DB07651; DB02662; DB08371; DB02977; DB06333; DB02436; DB04285; DB02651; DB03154 Interacts with Q13520; P56945; P11274-1; P07384; Q03135; Q14247; P00533; Q9GZR5; P19235; P10912; P62993; P08069; P06213; P06213-1; P05556; P05106; O60674; O43561; P08581; P04629; Q16288; P09619; P57054; P08922; P12931; P40763; P42229; Q9NPL8; Q96HV5; Q8N661; Q9H1D0; P10599; Q61140; Q63767; P15116; Q63768; P62994; P35570; P05622; P10686; P34152; Q8VI36; Q9WUD9; P63166 EC number EC 3.1.3.48 Uniprot keywords 3D-structure; Acetylation; Direct protein sequencing; Endoplasmic reticulum; Hydrolase; Membrane; Oxidation; Phosphoprotein; Protein phosphatase; Proteomics identification; Reference proteome; S-nitrosylation Protein physicochemical properties Chain ID A Molecular weight (Da) 34541 Length 297 Aromaticity 0.1 Instability index 35.91 Isoelectric point 5.91 Charge (pH=7) -5.37 3D Binding mode Sequence EMEKEFEQIDKSGSWAAIYQDIRHEASDFPCRVAKLPKNKNRNRYRDVSPFDHSRIKLHQEDNDYINASLIKMEEAQRSYILTQGPLPNTCGHFWEMVWEQKSRGVVMLNRVMEKGSLKCAQYWPQKEEKEMIFEDTNLKLTLISEDIKSYYTVRQLELENLTTQETREILHFHYTTWPDFGVPESPASFLNFLFKVRESGSLSPEHGPVVVHCSAGIGRSGTFCLADTCLLLMDKRKDPSSVDIKKVLLEMRKFRMGLIQTADQLRFSYLAVIEGAKFIMGDSSVQDQWKELSHED Hydrogen bonds contact Hydrophobic contact | ||||
| 20 | 2-oxoglutarate dehydrogenase, mitochondrial | 3ERY | 4.59 | |
Target general information Gen name OGDH Organism Homo sapiens (Human) Uniprot ID TTD ID NA Synonyms NA Protein family Alpha-ketoglutarate dehydrogenase family Biochemical class Immune system Function Chaperone binding.Heat shock protein binding.Metal ion binding.Oxoglutarate dehydrogenase (NAD+) activity.Oxoglutarate dehydrogenase (succinyl-transferring) activity.Thiamine pyrophosphate binding. Related diseases Postaxial acrofacial dysostosis (POADS) [MIM:263750]: POADS is characterized by severe micrognathia, cleft lip and/or palate, hypoplasia or aplasia of the posterior elements of the limbs, coloboma of the eyelids and supernumerary nipples. POADS is a very rare disorder: only 2 multiplex families, each consisting of 2 affected siblings born to unaffected, nonconsanguineous parents, have been described among a total of around 30 reported cases. {ECO:0000269|PubMed:19915526}. The disease is caused by variants affecting the gene represented in this entry. Drugs (DrugBank ID) DB00157; DB00313; DB09092 Interacts with P54253; P42858 EC number 1.2.4.2 Uniprot keywords 3D-structure; Acetylation; Alternative splicing; Calcium; Glycolysis; Isopeptide bond; Magnesium; Metal-binding; Mitochondrion; Nucleus; Oxidoreductase; Phosphoprotein; Proteomics identification; Reference proteome; Thiamine pyrophosphate; Transit peptide; Ubl conjugation Protein physicochemical properties Chain ID P,Q Molecular weight (Da) 18311.1 Length 158 Aromaticity 0.15 Instability index 37.26 Isoelectric point 5.64 Charge (pH=7) -2.98 3D Binding mode Sequence QLSPFPFDLGPHSMRYYETATSRRGLGEPRYTSVGYVDDKEFVRFDSDARITQVAKGQEQWFRVNLRTLLGYYNQSAGGTHTLQRMYGCDVGSDGRLLRGYEQFAYDGCDYIALNEDLRTWTAADMAAQITRRKWEQAGAAEYYRAYLEGECVEWLHR Hydrogen bonds contact Hydrophobic contact | ||||