Job Results:
Ligand
Structure
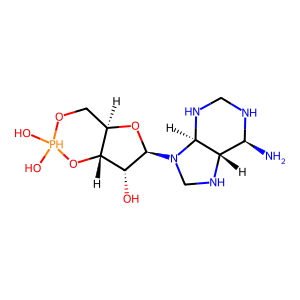
Job ID
e848ab91d8ae28235d6e66c6c8c71011
Job name
NA
Time
2024-09-30 12:19:50
| Rank | Target | PDB ID |
AirScore |
Detail |
|---|---|---|---|---|
| 61 | ATM serine/threonine kinase (ATM) | 7SIC | 6.52 | |
Target general information Gen name ATM Organism Homo sapiens (Human) Uniprot ID TTD ID Synonyms Serine-protein kinase ATM; Ataxia telangiectasia mutated; A-T mutated Protein family PI3/PI4-kinase family, ATM subfamily Biochemical class Kinase Function Serine/threonine protein kinase which activates checkpoint signaling upon double strand breaks (DSBs), apoptosis and genotoxic stresses such as ionizing ultraviolet A light (UVA), thereby acting as a DNA damage sensor. Recognizes the substrate consensus sequence [ST]-Q. Phosphorylates 'Ser-139' of histone variant H2AX/H2AFX at double strand breaks (DSBs), thereby regulating DNA damage response mechanism. Also plays a role in pre-B cell allelic exclusion, a process leading to expression of a single immunoglobulin heavy chain allele to enforce clonality and monospecific recognition by the B-cell antigen receptor (BCR) expressed on individual B-lymphocytes. After the introduction of DNA breaks by the RAG complex on one immunoglobulin allele, acts by mediating a repositioning of the second allele to pericentromeric heterochromatin, preventing accessibility to the RAG complex and recombination of the second allele. Also involved in signal transduction and cell cycle control. May function as a tumor suppressor. Necessary for activation of ABL1 and SAPK. Phosphorylates DYRK2, CHEK2, p53/TP53, FANCD2, NFKBIA, BRCA1, CTIP, nibrin (NBN), TERF1, RAD9, UBQLN4 and DCLRE1C. May play a role in vesicle and/or protein transport. Could play a role in T-cell development, gonad and neurological function. Plays a role in replication-dependent histone mRNA degradation. Binds DNA ends. Phosphorylation of DYRK2 in nucleus in response to genotoxic stress prevents its MDM2-mediated ubiquitination and subsequent proteasome degradation. Phosphorylates ATF2 which stimulates its function in DNA damage response. Related diseases Ataxia telangiectasia (AT) [MIM:208900]: A rare recessive disorder characterized by progressive cerebellar ataxia, dilation of the blood vessels in the conjunctiva and eyeballs, immunodeficiency, growth retardation and sexual immaturity. Patients have a strong predisposition to cancer; about 30% of patients develop tumors, particularly lymphomas and leukemias. Cells from affected individuals are highly sensitive to damage by ionizing radiation and resistant to inhibition of DNA synthesis following irradiation. {ECO:0000269|PubMed:10234507, ECO:0000269|PubMed:10425038, ECO:0000269|PubMed:10817650, ECO:0000269|PubMed:10873394, ECO:0000269|PubMed:19431188, ECO:0000269|PubMed:27664052, ECO:0000269|PubMed:7792600, ECO:0000269|PubMed:8589678, ECO:0000269|PubMed:8665503, ECO:0000269|PubMed:8698354, ECO:0000269|PubMed:8755918, ECO:0000269|PubMed:8789452, ECO:0000269|PubMed:8797579, ECO:0000269|PubMed:8808599, ECO:0000269|PubMed:8845835, ECO:0000269|PubMed:9043869, ECO:0000269|PubMed:9150358, ECO:0000269|PubMed:9443866, ECO:0000269|PubMed:9450874, ECO:0000269|PubMed:9463314, ECO:0000269|PubMed:9497252, ECO:0000269|PubMed:9521587, ECO:0000269|PubMed:9711876, ECO:0000269|PubMed:9792409, ECO:0000269|PubMed:9792410, ECO:0000269|PubMed:9872980, ECO:0000269|PubMed:9887333}. The disease is caused by variants affecting the gene represented in this entry.; DISEASE: Defects in ATM may contribute to T-cell acute lymphoblastic leukemia (TALL) and T-prolymphocytic leukemia (TPLL). TPLL is characterized by a high white blood cell count, with a predominance of prolymphocytes, marked splenomegaly, lymphadenopathy, skin lesions and serous effusion. The clinical course is highly aggressive, with poor response to chemotherapy and short survival time. TPLL occurs both in adults as a sporadic disease and in younger AT patients. {ECO:0000269|PubMed:9288106, ECO:0000269|PubMed:9334731, ECO:0000269|PubMed:9463314, ECO:0000269|PubMed:9488043, ECO:0000269|PubMed:9573030}.; DISEASE: Defects in ATM may contribute to B-cell non-Hodgkin lymphomas (BNHL), including mantle cell lymphoma (MCL). {ECO:0000269|PubMed:10397742, ECO:0000269|PubMed:10706620, ECO:0000269|PubMed:9288106}.; DISEASE: Defects in ATM may contribute to B-cell chronic lymphocytic leukemia (BCLL). BCLL is the commonest form of leukemia in the elderly. It is characterized by the accumulation of mature CD5+ B-lymphocytes, lymphadenopathy, immunodeficiency and bone marrow failure. {ECO:0000269|PubMed:10023947, ECO:0000269|PubMed:10397742, ECO:0000269|PubMed:9892178}. Drugs (DrugBank ID) DB00201 Interacts with Q9NY61; P00519; O43313; Q6PJG6; P62508-3; Q5XUX0; Q9Y6K9; Q13007; Q14676; Q9BQ15; P11245; O60934; P46531; Q9BZ95; Q7LG56; Q9Y4R8; P54274; P54274-2; Q15554; Q12888; O43156; PRO_0000037577 [P27958] EC number EC 2.7.11.1 Uniprot keywords 3D-structure; Acetylation; ATP-binding; Cell cycle; Cytoplasm; Cytoplasmic vesicle; Cytoskeleton; Disease variant; DNA damage; DNA-binding; Kinase; Neurodegeneration; Nucleotide-binding; Nucleus; Peroxisome; Phosphoprotein; Proteomics identification; Reference proteome; Serine/threonine-protein kinase; Transferase; Tumor suppressor Protein physicochemical properties Chain ID A Molecular weight (Da) 318958 Length 2773 Aromaticity 0.09 Instability index 42.93 Isoelectric point 7 Charge (pH=7) -0.06 3D Binding mode Sequence LNDLLICCRQLEHDRATERKKEVEKFKRLIRDPETIKHLDNWDAVFRFLQKYIQKETECQASRQKKMQEISSLVKYFIKCANRRAPRLKCQELLNYIMDTVKDSSNGAIYGADCSNILLKDILSVRKYWCEISQQQWLELFSVYFRLYLKPSQDVHRVLVARIIHAVTKGCCSQTDGLNSKFLDFFSKAIQCARQEKSSSGLNHILAALTIFLKTLAVNFRIRVCELGDEILPTLLYIWTQHRLNDSLKEVIIELFQLQIYIHHPKGAKTQEKGAYESTKWRSILYNLYDLLVNEISHIGSRRNIAVKENLIELMADICHQVFNIELGWEVIKDHLQKSQNDFDLVPWLQIATQLISKYPASLPNCELSPLLMILSQLLPQQRHGERTPYVLRCLTEVALCQDKRSNLESSQKSDLLKLWNKIWCITFRGISSEQIQAENFGLLGAIIQGSLVEVDREFWKLFTGSACRPSCPAVCCLTLALTTSIVPGTFSLKESIMKWLLFYQLETEVPPILHSNFPHLVLEKILVSLTMKNCKAAMNFFQSVPECSFSEVEELFLQTTFDKMDFLTGFSVHQNLKESLDRCLLGLSEQLLNNYSSEITNSETLVRCSRLLVGVLGCYCYMGVIAEEEAYKSELFQKAKSLMQCAGESITLFKNKTNEEFRIGSLRNMMQLCTRCLSNCTKKSPNKIASGFFLRLLTSKLMNDIADICKSLASFISTIGAINPLAEEYLSKQDLLFLDMLKFLCLCVTTAQTNTVSFRAADIRRKLLMLIDSSTLEPTKSLHLHMYLMLLKELPGEEYPLPMEDVLELLKPLSNVCSLYRRDQDVCKTILNHVLHVVKNLGQSNMDSENTRDAQGQFLTVIGAFWHLTKERKYIFSVRMALVNCLKTLLEADPYSKWAILNVMGKDFPVNEVFTQFLADNHHQVRMLAAESINRLFQDTKLKALPLKLQQTAFENAYLKAQEGMREMETLDEIYNRKSVLLTLIAVVLSCSPICEKQALFALCKSVKENGLEPHLVKKVLEKVSETFGYRRLEDFMASHLDYLVLEWLNLQDTEYNLSSFPFILLNYTNIEDFYRSCYKVLIPHLVIRSHFDEVKSIANQIQEDWKSLLTDCFPKILVNILPYFAYEGTRDSGMAQQRETATKVYDMLKSENLLGKQIDHLFISNLPEIVVELLMTLHEPDLDPAPNPPHFPSHVIKATFAYISNCHKTKLKSILEILSKSPDSYQKILLAICEQAAETNNVYKKHRILKIYHLFVSLLLKDIKSGLGGAWAFVLRDVIYTLIHYINQRPSCIMDVSLRSFSLCCDLLSQVCQTAVTYCKDALENHLHVIVGTLIPLVYEQVEVQKQVLDLLKYLVIDNKDNENLYITIKLLDPFPDHVVFKDLRITQQKIKYSRGPFSLLEEINHFLSVSVYDALPLTRLEGLKDLRRQLELHKDQMVDIMRASQDNPQDGIMVKLVVNLLQLSKMAINHTGEKEVLEAVGSCLGEVGPIDFSTIAIQHSKDASYTKALKLFEDKELQWTFIMLTYLNNTLVEDCVKVRSAAVTCLKNILATKTGHSFWEIYKMTTDPMLAYLQPFRTSRKKFLEVPRFDKENPFEGLDDINLWIPLSENHDIWIKTLTCAFLDSGGTKCEILQLLKPMCEVKTDFCQTVLPYLIHDILLQDTNESWRNLLSTHVQGFFTSCLRHCCLDKKSQRTMLAVVDYMRRQKRPSSGTIFNDAFWLDLNYLEVAKVAQSCAAHFTALLYAEIYADKKSMDDQEKRSTTISSLSEKSKEETGISLQDLLLEIYRSIGEPDSLYGCGGGKMLQPITRLRTYEHEAMWGKALVTYDLETAIPSSTRQAGIIQALQNLGLCHILSVYLKGLDYENKDWCPELEELHYQAAWRNMQWDHCGTSYHESLYNALQSLRDREFSTFYESLKYARVKEVEEMCKRSLESVYSLYPTLSRLQAIGELESIGELFSRSVTHRQLSEVYIKWQKHSQLLKDSDFSFQEPIMALRTVILEILMEKEMDNSQRECIKDILTKHLVELSILARTFKNTQLPERAIFQIKQYNSVSCGVSEWQLEEAQVFWAKKEQSLALSILKQMIKKLDASCAANNPSLKLTYTECLRVCGNWLAETCLENPAVIMQTYLEKAVEVAGNYDGESSDELRNGKMKAFLSLARFSDTQYQRIENYMKSSEFENKQALLKRAKERYTVKVQRELELDELALRALKEDRKRFLCKAVENYINCLLSGEEHDMWVFRLCSLWLENSGVSEVNGMMKRDGMKIPTYKFLPLMYQLAARMGTKMMGGLGFHEVLNNLISRISMDHPHHTLFIILALANANRDEFLTSSQLDEDRTEAANRIICTIRSRRPQMVRSVEALCDAYIILANLDATQWKTQRKGINIPADQPITKLKNLEDVVVPTMEIKVDHTGEYGNLVTIQSFKAEFRLAGGVNLPKIIDCVGSDGKERRQLVKGRDDLRQDAVMQQVFQMCNTLLQRNTETRKRKLTICTYKVVPLSQRSGVLEWCTGTVPIGEFLVNNEDGAHKRYRPNDFSAFQCQKKMMEVQKKSFEEKYEVFMDVCQNFQPVFRYFCMEKFLDPAIWFEKRLAYTRSVATSSIVGYILGLGDRHVQNILINEQSAELVHIDLGVAFEQGKILPTPETVPFRLTRDIVDGMGITGVEGVFRRCCEKTMEVMRNSQETLLTIVEVLLYDPLFDWTMNPLKALYLQQRPSFNKVAERVLMRLQEKLKGVEEGTVLSVGGQVNLLIQQAIDPKNLSRLFPGWKAWV Hydrogen bonds contact Hydrophobic contact | ||||
| 62 | Fungal Isoleucyl t-RNA synthetase (Fung ILS1) | 7D5C | 6.51 | |
Target general information Gen name Fung ILS1 Organism Saccharomyces cerevisiae (strain ATCC 204508 / S288c) (Baker's yeast) Uniprot ID TTD ID Synonyms Fung Isoleucyl-tRNA synthetase; Fung Isoleucine--tRNA ligase, cytoplasmic; Fung IleRS Protein family Class-I aminoacyl-tRNA synthetase family Biochemical class Carbon-oxygen ligase Function Has aminoacyl-tRNA editing and isoleucine-tRNA ligase activity. Related diseases Intellectual developmental disorder with dysmorphic facies and ptosis (IDDDFP) [MIM:617333]: An autosomal dominant neurodevelopmental disorder characterized by delayed psychomotor development, intellectual disability, delayed language, and facial dysmorphisms, most notably ptosis. Additional features may include poor growth, hypotonia, and seizures. {ECO:0000269|PubMed:27939639, ECO:0000269|PubMed:27939640}. The disease is caused by variants affecting the gene represented in this entry. Drugs (DrugBank ID) NA Interacts with NA EC number EC 6.1.1.5 Uniprot keywords 3D-structure; Aminoacyl-tRNA synthetase; ATP-binding; Cytoplasm; Direct protein sequencing; Isopeptide bond; Ligase; Nucleotide-binding; Phosphoprotein; Protein biosynthesis; Reference proteome; Ubl conjugation Protein physicochemical properties Chain ID A Molecular weight (Da) 105860 Length 919 Aromaticity 0.12 Instability index 37.01 Isoelectric point 5.73 Charge (pH=7) -14.6 3D Binding mode Sequence FSFPKEEEKVLSLWDEIDAFHTSLELTKDKPEFSFFDGPPFATGTPHYGHILASTIKDIVPRYATMTGHHVERRFGWDTHGVPIEHIIDKKLGITGKDDVFKYGLENYNNECRSIVMTYASDWRKTIGRLGRWIDFDNDYKTMYPSFMESTWWAFKQLHEKGQVYRGFKVMPYSTGLTTPLSNFEAQQNYKDVNDPAVTIGFNVIGQEKTQLVAWTTTPWTLPSNLSLCVNADFEYVKIYDETRDRYFILLESLIKTLYKKPKNEKYKIVEKIKGSDLVGLKYEPLFPYFAEQFHETAFRVISDDYVTSDSGTGIVHNAPAFGEEDNAACLKNGVISEDSVLPNAIDDLGRFTKDVPDFEGVYVKDADKLIIKYLTNTGNLLLASQIRHSYPFCWRSDTPLLYRSVPAWFVRVKNIVPQMLDSVMKSHWVPNTIKEKRFANWIANARDWNVSRNRYWGTPIPLWVSDDFEEVVCVGSIKELEELTGVRNITDLHRDVIDKLTIPSKQGKGDLKRIEEVFDCWFESGSMPYASQHYPFENTEKFDERVPANFISEGLDQTRGWFYTLAVLGTHLFGSVPYKNVIVSGIVLAADGRKMSKSLKNYPDPSIVLNKYGADALRLYLINSPVLKAESLKFKEEGVKEVVSKVLLPWWNSFKFLDGQIALLKKMSNIDFQYDDSVKSDNVMDRWILASMQSLVQFIHEEMGQYKLYTVVPKLLNFIDELTNWYIRFNRRRLKGENGVEDCLKALNSLFDALFTFVRAMAPFTPFLSESIYLRLKEYIPEAVLAKYGKDGRSVHFLSYPVVKKEYFDEAIETAVSRMQSVIDLGRNIREKKTISLKTPLKTLVILHSDESYLKDVEALKNYIIEELNVRDVVITSDEAKYGVEYRGLPESAVQAGQETRTDQDVLIIMDTNIYSEL Hydrogen bonds contact Hydrophobic contact | ||||
| 63 | Ribosomal protein S6 kinase beta-1 (S6K1) | 7N93 | 6.51 | |
Target general information Gen name RPS6KB1 Organism Homo sapiens (Human) Uniprot ID TTD ID Synonyms p70-S6K 1; p70 ribosomal S6 kinase alpha; p70 S6KA; p70 S6K-alpha; p70 S6 kinase alpha; Serine/threonine-protein kinase 14A; STK14A; S6K-beta-1; S6K; Ribosomal protein S6 kinase I; P70S6K1; P70-S6K; 7 Protein family Protein kinase superfamily, AGC Ser/Thr protein kinase family, S6 kinase subfamily Biochemical class Kinase Function Regulates protein synthesis through phosphorylation of EIF4B, RPS6 and EEF2K, and contributes to cell survival by repressing the pro-apoptotic function of BAD. Under conditions of nutrient depletion, the inactive form associates with the EIF3 translation initiation complex. Upon mitogenic stimulation, phosphorylation by the mammalian target of rapamycin complex 1 (mTORC1) leads to dissociation from the EIF3 complex and activation. The active form then phosphorylates and activates several substrates in the pre-initiation complex, including the EIF2B complex and the cap-binding complex component EIF4B. Also controls translation initiation by phosphorylating a negative regulator of EIF4A, PDCD4, targeting it for ubiquitination and subsequent proteolysis. Promotes initiation of the pioneer round of protein synthesis by phosphorylating POLDIP3/SKAR. In response to IGF1, activates translation elongation by phosphorylating EEF2 kinase (EEF2K), which leads to its inhibition and thus activation of EEF2. Also plays a role in feedback regulation of mTORC2 by mTORC1 by phosphorylating RICTOR, resulting in the inhibition of mTORC2 and AKT1 signaling. Mediates cell survival by phosphorylating the pro-apoptotic protein BAD and suppressing its pro-apoptotic function. Phosphorylates mitochondrial URI1 leading to dissociation of a URI1-PPP1CC complex. The free mitochondrial PPP1CC can then dephosphorylate RPS6KB1 at Thr-412, which is proposed to be a negative feedback mechanism for the RPS6KB1 anti-apoptotic function. Mediates TNF-alpha-induced insulin resistance by phosphorylating IRS1 at multiple serine residues, resulting in accelerated degradation of IRS1. In cells lacking functional TSC1-2 complex, constitutively phosphorylates and inhibits GSK3B. May be involved in cytoskeletal rearrangement through binding to neurabin. Phosphorylates and activates the pyrimidine biosynthesis enzyme CAD, downstream of MTOR. Following activation by mTORC1, phosphorylates EPRS and thereby plays a key role in fatty acid uptake by adipocytes and also most probably in interferon-gamma-induced translation inhibition. Serine/threonine-protein kinase that acts downstream of mTOR signaling in response to growth factors and nutrients to promote cell proliferation, cell growth and cell cycle progression. Related diseases Progressive familial heart block 1B (PFHB1B) [MIM:604559]: A cardiac bundle branch disorder characterized by progressive alteration of cardiac conduction through the His-Purkinje system, with a pattern of a right bundle-branch block and/or left anterior hemiblock occurring individually or together. It leads to complete atrio-ventricular block causing syncope and sudden death. {ECO:0000269|PubMed:19726882, ECO:0000269|PubMed:20562447, ECO:0000269|PubMed:21887725}. The disease is caused by variants affecting the gene represented in this entry.; DISEASE: Erythrokeratodermia variabilis et progressiva 6 (EKVP6) [MIM:618531]: A form of erythrokeratodermia variabilis et progressiva, a genodermatosis characterized by the coexistence of two independent skin lesions: transient erythema and hyperkeratosis that is usually localized but occasionally occurs in its generalized form. Clinical presentation varies significantly within a family and from one family to another. Palmoplantar keratoderma is present in around 50% of cases. EKVP6 inheritance is autosomal dominant. {ECO:0000269|PubMed:30528822}. The disease is caused by variants affecting the gene represented in this entry. Drugs (DrugBank ID) NA Interacts with P31749; Q06481-5; P55884; P08151; Q5VY09; Q00005; P13051-2; P08151; P05067; P27986-2 EC number EC 2.7.11.1 Uniprot keywords 3D-structure; Acetylation; Alternative initiation; Alternative splicing; Apoptosis; ATP-binding; Cell cycle; Cytoplasm; Kinase; Membrane; Mitochondrion; Mitochondrion outer membrane; Nucleotide-binding; Nucleus; Phosphoprotein; Proteomics identification; Reference proteome; Serine/threonine-protein kinase; Synapse; Synaptosome; Transferase; Translation regulation Protein physicochemical properties Chain ID A Molecular weight (Da) 35187.6 Length 312 Aromaticity 0.1 Instability index 39.58 Isoelectric point 8.8 Charge (pH=7) 5.54 3D Binding mode Sequence RPECFELLRVLGKGGYGKVFQVRKVTGANTGKIFAMKVLKKAMIVRNAKDTAHTKAERNILEEVKHPFIVDLIYAFQTGGKLYLILEYLSGGELFMQLEREGIFMEDTACFYLAEISMALGHLHQKGIIYRDLKPENIMLNHQGHVKLTDFGLCKECGTIEYMAPEILMRSGHNRAVDWWSLGALMYDMLTGAPPFTGENRKKTIDKILKCKLNLPPYLTQEARDLLKKLLKRNAASRLGAGPGDAGEVQAHPFFRHINWEELLARKVEPPFKPLLQSEEDVSQFDSKFTRQXPVDXPNQVFLGFEYVAPSV Hydrogen bonds contact Hydrophobic contact | ||||
| 64 | Phosphodiesterase 10A (PDE10) | 2OUR | 6.50 | |
Target general information Gen name PDE10A Organism Homo sapiens (Human) Uniprot ID TTD ID Synonyms cAMP and cAMPinhibited cGMP 3',5'cyclic phosphodiesterase 10A; cAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A Protein family Cyclic nucleotide phosphodiesterase family Biochemical class Phosphoric diester hydrolase Function Can hydrolyze both cAMP and cGMP, but has higher affinity for cAMP and is more efficient with cAMP as substrate. May play a critical role in regulating cAMP and cGMP levels in the striatum, a region of the brain that contributes to the control of movement and cognition. Plays a role in signal transduction by regulating the intracellular concentration of cyclic nucleotides. Related diseases Dyskinesia, limb and orofacial, infantile-onset (IOLOD) [MIM:616921]: An autosomal recessive, early-onset hyperkinetic movement disorder characterized by axial hypotonia, dyskinesia of the limbs and trunk, orofacial dyskinesia, drooling, and dysarthria. The severity of the hyperkinesis is variable. {ECO:0000269|PubMed:27058446}. The disease is caused by variants affecting the gene represented in this entry.; DISEASE: Striatal degeneration, autosomal dominant 2 (ADSD2) [MIM:616922]: An autosomal dominant disorder characterized by striatal degeneration and dysfunction of basal ganglia, resulting in hyperkinesis. {ECO:0000269|PubMed:27058447}. The disease is caused by variants affecting the gene represented in this entry. Drugs (DrugBank ID) DB08384; DB08386; DB08383; DB08389; DB00201; DB00975; DB08387; DB01113; DB08391; DB08811; DB09283; DB08814 Interacts with NA EC number EC 3.1.4.17 Uniprot keywords 3D-structure; Allosteric enzyme; Alternative initiation; Alternative splicing; cAMP; cAMP-binding; cGMP; cGMP-binding; Cytoplasm; Disease variant; Hydrolase; Metal-binding; Nucleotide-binding; Phosphoprotein; Proteomics identification; Reference proteome; Repeat Protein physicochemical properties Chain ID A,B Molecular weight (Da) 37469.9 Length 324 Aromaticity 0.1 Instability index 48.52 Isoelectric point 6.28 Charge (pH=7) -4.08 3D Binding mode Sequence HMSICTSEEWQGLMQFTLPVRLCKEIELFHFDIGPFENMWPGIFVYMVHRSCGTSCFELEKLCRFIMSVKKNYRRVPYHNWKHAVTVAHCMYAILQNNHTLFTDLERKGLLIACLCHDLDHRGFSNSYLQKFDHPLAALYSTSTMEQHHFSQTVSILQLEGHNIFSTLSSSEYEQVLEIIRKAIIATDLALYFGNRKQLEEMYQTGSLNLNNQSHRDRVIGLMMTACALCSVTKLWPVTKLTANDIYAEFWAEGDEMKKLGIQPIPMMDRDKKDEVPQGQLGFYNAVAIPCYTTLTQILPPTEPLLKACRDNLSQWEKVIRGEE Hydrogen bonds contact Hydrophobic contact | ||||
| 65 | Geranylgeranyl pyrophosphate synthase | 2Q80 | 6.50 | |
Target general information Gen name GGPS1 Organism Homo sapiens (Human) Uniprot ID TTD ID NA Synonyms NA Protein family FPP/GGPP synthase family Biochemical class Transferase Function Dimethylallyltranstransferase activity.Farnesyltranstransferase activity.Geranyltranstransferase activity.Identical protein binding.Metal ion binding. Related diseases Muscular dystrophy, congenital hearing loss, and ovarian insufficiency syndrome (MDHLO) [MIM:619518]: An autosomal recessive disorder characterized by early-onset progressive muscle weakness, sensorineural hearing loss, and primary amenorrhea due to ovarian insufficiency. Some patients become wheelchair-bound by the second decade, whereas others have a milder phenotype and maintain independent ambulation into adulthood. Most patients have respiratory insufficiency. {ECO:0000269|PubMed:32403198}. The disease is caused by variants affecting the gene represented in this entry. Drugs (DrugBank ID) DB06830; DB06931; DB07221; DB08529; DB07410; DB07780; DB04695; DB02552; DB07841; DB00710; DB04714; DB07873; DB06548; DB00282; DB00399 Interacts with O00244; O95749; O00560 EC number 2.5.1.-; 2.5.1.1; 2.5.1.10; 2.5.1.29 Uniprot keywords 3D-structure; Acetylation; Alternative splicing; Congenital muscular dystrophy; Cytoplasm; Deafness; Disease variant; Isoprene biosynthesis; Lipid metabolism; Magnesium; Metal-binding; Proteomics identification; Reference proteome; Transferase Protein physicochemical properties Chain ID A,B,C,D,E,F Molecular weight (Da) 32872.4 Length 284 Aromaticity 0.11 Instability index 41.09 Isoelectric point 6.17 Charge (pH=7) -3.37 3D Binding mode Sequence ETVQRILLEPYKYLLQLPGKQVRTKLSQAFNHWLKVPEDKLQIIIEVTEMLHNASLLIDDIEDNSKLRRGFPVAHSIYGIPSVINSANYVYFLGLEKVLTLDHPDAVKLFTRQLLELHQGQGLDIYWRDNYTCPTEEEYKAMVLQKTGGLFGLAVGLMQLFSDYKEDLKPLLNTLGLFFQIRDDYANLHSKSFCEDLTEGKFSFPTIHAIWSRPESTQVQNILRQRTENIDIKKYCVHYLEDVGSFEYTRNTLKELEAKAYKQIDARGGNPELVALVKHLSKMF Hydrogen bonds contact Hydrophobic contact | ||||
| 66 | Wnt-7a protein (WNT7A) | 4UZQ | 6.49 | |
Target general information Gen name WNT7A Organism Homo sapiens (Human) Uniprot ID TTD ID Synonyms Protein Wnt-7a Protein family Wnt family Biochemical class NA Function Plays an important role in embryonic development, including dorsal versus ventral patterning during limb development, skeleton development and urogenital tract development. Required for central nervous system (CNS) angiogenesis and blood-brain barrier regulation. Required for normal, sexually dimorphic development of the Mullerian ducts, and for normal fertility in both sexes. Required for normal neural stem cell proliferation in the hippocampus dentate gyrus. Required for normal progress through the cell cycle in neural progenitor cells, for self-renewal of neural stem cells, and for normal neuronal differentiation and maturation. Promotes formation of synapses via its interaction with FZD5. Ligand for members of the frizzled family of seven transmembrane receptors that functions in the canonical Wnt/beta-catenin signaling pathway. Related diseases Limb pelvis hypoplasia aplasia syndrome (LPHAS) [MIM:276820]: A syndrome of severe deficiency of the extremities due to hypo- or aplasia of one or more long bones of one or more limbs. Pelvic manifestations include hip dislocation, hypoplastic iliac bone and aplastic pubic bones. Thoracic deformity, unusual facies and genitourinary anomalies can be present. {ECO:0000269|PubMed:16826533, ECO:0000269|PubMed:17431918, ECO:0000269|PubMed:20949531, ECO:0000269|PubMed:21271649, ECO:0000269|PubMed:27638328}. The disease is caused by variants affecting the gene represented in this entry.; DISEASE: Fuhrmann syndrome (FUHRS) [MIM:228930]: Distinct limb-malformation disorder characterized also by various degrees of limb aplasia/hypoplasia and joint dysplasia. {ECO:0000269|PubMed:16826533}. The disease is caused by variants affecting the gene represented in this entry. Drugs (DrugBank ID) NA Interacts with P55212; P22607; P06396; P13473-2; Q9UMX0; Q9Y5W5; Q5T9L3; Q9Z0J1 EC number NA Uniprot keywords 3D-structure; Developmental protein; Disease variant; Disulfide bond; Extracellular matrix; Glycoprotein; Lipoprotein; Proteomics identification; Reference proteome; Secreted; Signal; Wnt signaling pathway Protein physicochemical properties Chain ID A,B Molecular weight (Da) 40475.5 Length 356 Aromaticity 0.1 Instability index 50.49 Isoelectric point 7.67 Charge (pH=7) 1.62 3D Binding mode Sequence EDLRLHLLLNTSVTCNDGSPAGYYLKESRGSRRWLLFLEGGWYCFNRENCDSRYDTMRRLMSSRDWPRTRTGTGILSSQPEENPYWWNANMVFIPYCSSDVWSGASSKSEKNEYAFMGALIIQEVVRELLGRGLSGAKVLLLAGSAAGGTGVLLNVDRVAEQLEKLGYPAIQVRGLADSGWFLDNKQYRHTDCVDTITCAPTEAIRRGIRYWNGVVPERCRRQFQEGEEWNCFFGYKVYPTLRSPVFVVQWLFDEAQLTVDNVHLTGQPVQEGLRLYIQNLGRELRHTLKDVPASFAPACLSHEIIIRSHWTDVQVKGTSLPRALHCWDRSLHKGCPVHLVDSCPWPHCNPSCPTS Hydrogen bonds contact Hydrophobic contact | ||||
| 67 | Caspase-6 (CASP6) | 4NBL | 6.48 | |
Target general information Gen name CASP6 Organism Homo sapiens (Human) Uniprot ID TTD ID Synonyms MCH2; Caspase-6 subunit p18; Caspase-6 subunit p11; CASP-6; Apoptotic protease Mch-2 Protein family Peptidase C14A family Biochemical class Peptidase Function Cleaves poly(ADP-ribose) polymerase in vitro, as well as lamins. Overexpression promotes programmed cell death. Involved in the activation cascade of caspases responsible for apoptosis execution. Related diseases Growth hormone deficiency, isolated, 1A (IGHD1A) [MIM:262400]: An autosomal recessive, severe deficiency of growth hormone leading to dwarfism. Patients often develop antibodies to administered growth hormone. {ECO:0000269|PubMed:8364549}. The disease is caused by variants affecting the gene represented in this entry.; DISEASE: Growth hormone deficiency, isolated, 1B (IGHD1B) [MIM:612781]: An autosomal recessive deficiency of growth hormone leading to short stature. Patients have low but detectable levels of growth hormone, significantly retarded bone age, and a positive response and immunologic tolerance to growth hormone therapy. {ECO:0000269|PubMed:12655557}. The disease is caused by variants affecting the gene represented in this entry.; DISEASE: Kowarski syndrome (KWKS) [MIM:262650]: A syndrome clinically characterized by short stature associated with bioinactive growth hormone, normal or slightly increased growth hormone secretion, pathologically low insulin-like growth factor 1 levels, and normal catch-up growth on growth hormone replacement therapy. {ECO:0000269|PubMed:17519310, ECO:0000269|PubMed:8552145, ECO:0000269|PubMed:9276733}. The disease is caused by variants affecting the gene represented in this entry.; DISEASE: Growth hormone deficiency, isolated, 2 (IGHD2) [MIM:173100]: An autosomal dominant deficiency of growth hormone leading to short stature. Clinical severity is variable. Patients have a positive response and immunologic tolerance to growth hormone therapy. {ECO:0000269|PubMed:11502836, ECO:0000269|PubMed:9152628}. The disease is caused by variants affecting the gene represented in this entry. Drugs (DrugBank ID) NA Interacts with Q9Y614; Q6DHV7-2; Q6UY14-3; Q96MA6; Q5T2L2; Q96Q83-2; Q9Y303-2; Q9NU02; P09525; P06727; Q8WW27; Q66PJ3-4; Q6XD76; P18848; Q9H0Y0; Q14032; P54687-4; P06276; Q9NSI6-4; Q96Q07-2; Q9H0W9-3; Q9NQ89; Q13901; Q3SXR2; Q8N1A6; P17655; P20807-4; P42574; P55212; O00257-3; P24863; Q9NNX6-10; Q9UJX2; P42773; O95674; Q494V2-2; Q8WUX9; Q9Y3D0; Q8N365; Q3SX64; Q99966; P09496-2; Q6PJW8-3; Q96BR5; P02458-1; Q9UGL9; Q9UKG9-2; P26998; P35222; Q53TN4; P61962; O60479; Q96EY1-3; Q92782-2; Q9BPU6; A0AVK6; Q658K8; O00303; Q13347; O00472; O00423; Q6NXG1-3; Q49AJ0-4; Q8N128-2; Q8IZU1; Q6ZNL6; Q9NSA1; Q06547-3; Q49A26-4; Q9HAV0; Q6NXT2; Q9BT25; Q9NRZ9-6; Q96EW2-2; P42858; Q8N6M8-2; Q92613; P0C870; Q9UK76; Q8N5Z5; Q8TBB5-2; Q9UH77; Q8N4N3-2; Q5JUW0-3; Q8N1A0; P13473-2; Q6DKI2; Q9H2C1; Q8N0U6; Q9Y234; Q8TBB1; Q1L5Z9; Q96JB6; Q16609; Q8IYG6; P0DP58-2; Q969L2; P27338; A6NJ78-4; Q96C03-3; Q8N5J2-3; A0A0A0MR05; P34949-2; Q9BV20; Q6IN84-2; A2RUH7; P01106; Q9H7X0; Q15742-2; Q9UJ70-2; Q8NDH3-5; Q96HA8; P36639-4; Q8NFH4; Q8NFH3; Q7Z3B4; Q6N063-2; Q6GQQ9-2; Q9H8K7; Q99447; P27815-4; O15534; Q9BUL5; Q00169; P48739; P61925; Q58EX7-2; O60664; Q14181; P0DPB6; P36954; Q07869; O60927; Q6ZMI0-5; P54619; Q8NCQ7-2; P41222; P29074; Q8WUD1-2; Q5R372-9; Q9HD47-3; Q09028; Q04206; P47804-3; Q15382; Q06587; Q8N5U6; P62701; Q66K80; Q01826; O15126; P22307-3; Q9BRK5; Q9NTN9-3; P01011; Q15393; Q9NR46; Q9BZQ2; O60902-3; Q86US8; P37840; Q96H20; Q13573; Q7Z6I5; Q496A3; Q9C004; Q5W111-2; Q96BD6; Q92797-2; O60506-4; O15273; Q86WV5; Q96A09; P54274-2; P22735; O43548; Q9NQ88; Q9UIK5-2; Q53NU3; P04637; Q12888; P36406; Q86WT6-2; Q13885; P49459; Q9P1Q0-4; Q9NX94; Q8NA23-2; Q9BQA1; O00755; O95070; O43829; Q8IWT0-2; Q53FD0-2; Q05CR2; Q96JL9-2; Q96LX8; Q3KNS6-3; Q5JTY5; A0A384MDV8; B7Z3E8; Q86V28 EC number EC 3.4.22.59 Uniprot keywords 3D-structure; Alternative splicing; Apoptosis; Autocatalytic cleavage; Cytoplasm; Hydrolase; Lipoprotein; Nucleus; Palmitate; Phosphoprotein; Protease; Proteomics identification; Reference proteome; Thiol protease; Zymogen Protein physicochemical properties Chain ID A,B Molecular weight (Da) 57170 Length 500 Aromaticity 0.12 Instability index 31.33 Isoelectric point 8.05 Charge (pH=7) 4.52 3D Binding mode Sequence AFYKREMFDPAEKYKMDHRRRGIALIFNHERFFWHLTLPERRGTCADRDNLTRRFSDLGFEVKCFNDLKAEELLLKIHEVSTVSHADADCFVCVFLSHGEGNHIYAYDAKIEIQTLTGLFKGDKCHSLVGKPKIFIIQAARGNQHDVPVIPDTNITEVDAASVYTLPAGADFLMCYSVAEGYYSHRETVNGSWYIQDLCEMLGKYGSSLEFTELLTLVNRKVSQRRVDFCKDPSAIGKKQVPCFASMLTKKLHFFPKSMFDPAEKYKMDHRRRGIALIFNHERFFWHLTLPERRGTCADRDNLTRRFSDLGFEVKCFNDLKAEELLLKIHEVSTVSHADADCFVCVFLSHGEGNHIYAYDAKIEIQTLTGLFKGDKCHSLVGKPKIFIIQAARGNTNITEVDAASVYTLPAGADFLMCYSVAEGYYSHRETVNGSWYIQDLCEMLGKYGSSLEFTELLTLVNRKVSQRRVDFCKDPSAIGKKQVPCFASMLTKKLHFFPK Hydrogen bonds contact Hydrophobic contact | ||||
| 68 | Suppressor of tumorigenicity 14 protein (ST14) | 3P8G | 6.48 | |
Target general information Gen name ST14 Organism Homo sapiens (Human) Uniprot ID TTD ID Synonyms Tumor-associated differentially-expressed gene 15 protein; Tumor associated differentially-expressed gene-15 protein; TADG15; Serine protease TADG-15; Serine protease 14; SNC19; Prostamin; PRSS14; Mem Protein family Peptidase S1 family Biochemical class Peptidase Function Proposed to play a role in breast cancer invasion and metastasis. Exhibits trypsin-like activity as defined by cleavage of synthetic substrates with Arg or Lys as the P1 site. Involved in the terminal differentiation of keratinocytes through prostasin (PRSS8) activation and filaggrin (FLG) processing. Degrades extracellular matrix. Related diseases Ichthyosis, congenital, autosomal recessive 11 (ARCI11) [MIM:602400]: A form of autosomal recessive congenital ichthyosis, a disorder of keratinization with abnormal differentiation and desquamation of the epidermis, resulting in abnormal skin scaling over the whole body. The main skin phenotypes are lamellar ichthyosis (LI) and non-bullous congenital ichthyosiform erythroderma (NCIE), although phenotypic overlap within the same patient or among patients from the same family can occur. Lamellar ichthyosis is a condition often associated with an embedment in a collodion-like membrane at birth; skin scales later develop, covering the entire body surface. Non-bullous congenital ichthyosiform erythroderma characterized by fine whitish scaling on an erythrodermal background; larger brownish scales are present on the buttocks, neck and legs. {ECO:0000269|PubMed:17273967, ECO:0000269|PubMed:18843291}. The disease is caused by variants affecting the gene represented in this entry. Drugs (DrugBank ID) DB03127; DB13729; DB00013 Interacts with NA EC number EC 3.4.21.109 Uniprot keywords 3D-structure; Disease variant; Disulfide bond; Glycoprotein; Hydrolase; Hypotrichosis; Ichthyosis; Membrane; Protease; Proteomics identification; Reference proteome; Repeat; Serine protease; Signal-anchor; Transmembrane; Transmembrane helix Protein physicochemical properties Chain ID A Molecular weight (Da) 26451.5 Length 241 Aromaticity 0.1 Instability index 30.45 Isoelectric point 5.6 Charge (pH=7) -5.69 3D Binding mode Sequence VVGGTDADEGEWPWQVSLHALGQGHICGASLISPNWLVSAAHCYIDDRGFRYSDPTQWTAFLGLHDQSQRSAPGVQERRLKRIISHPFFNDFTFDYDIALLELEKPAEYSSMVRPICLPDASHVFPAGKAIWVTGWGHTQYGGTGALILQKGEIRVIQQTTCENLLPQQITPRMMCVGFLSGGVDSCQGDSGGPLSSVEADGRIFQAGVVSWGDGCAQRNKPGVYTRLPLFRDWIKENTGV Hydrogen bonds contact Hydrophobic contact | ||||
| 69 | Organic cation transporter 3 (OCT3) | 7ZH6 | 6.47 | |
Target general information Gen name SLC22A3 Organism Homo sapiens (Human) Uniprot ID TTD ID Synonyms Solute carrier family 22 member 3; Extraneuronal monoamine transporter; EMTH Protein family Major facilitator (TC 2.A.1) superfamily, Organic cation transporter (TC 2.A.1.19) family Biochemical class NA Function Mediates potential-dependent transport of a variety of organic cations. May play a significant role in the disposition of cationic neurotoxins and neurotransmitters in the brain. Related diseases Deafness, autosomal dominant, 2A (DFNA2A) [MIM:600101]: A form of non-syndromic sensorineural hearing loss. Sensorineural deafness results from damage to the neural receptors of the inner ear, the nerve pathways to the brain, or the area of the brain that receives sound information. {ECO:0000269|PubMed:10025409, ECO:0000269|PubMed:10369879, ECO:0000269|PubMed:10571947, ECO:0000269|PubMed:10925378, ECO:0000269|PubMed:21242547}. The disease is caused by variants affecting the gene represented in this entry. Drugs (DrugBank ID) DB00718; DB08838; DB00182; DB00122; DB14006; DB00501; DB00575; DB00363; DB01151; DB00988; DB00783; DB13952; DB13953; DB13954; DB13955; DB13956; DB00983; DB00536; DB05381; DB00458; DB00762; DB00709; DB00448; DB08882; DB01042; DB01577; DB00331; DB08893; DB00184; DB00368; DB00526; DB00925; DB00413; DB00457; DB01035; DB00396; DB00938; DB00391; DB13943; DB13944; DB08837; DB08841; DB00541 Interacts with P00519 EC number NA Uniprot keywords 3D-structure; Cell membrane; Glycoprotein; Ion transport; Membrane; Mitochondrion; Nucleus; Proteomics identification; Reference proteome; Transmembrane; Transmembrane helix; Transport Protein physicochemical properties Chain ID A Molecular weight (Da) 53067.4 Length 478 Aromaticity 0.13 Instability index 38.82 Isoelectric point 9.07 Charge (pH=7) 10.54 3D Binding mode Sequence SFDEALQRVGEFGRFQRRVFLLLCLTGVTFAFLFVGVVFLGTQPDHYWCRGPSAAALAERCGWSPEEEWNRTAPASRGRCQRYLLSAPLVPCRGGWRYAQAHSTIVSEFDLVCVNAWMLDLTQAILNLGFLTGAFTLGYAADRYGRIVIYLLSCLGVGVTGVVVAFAPNFPVFVIFRFLQGVFGKGTWMTCYVIVTEIVGSKQRRIVGIVIQMFFTLGIIILPGIAYFIPNWQGIQLAITLPSFLFLLYYWVVPESPRWLITRKKGDKALQILRRIAKCNVSNPSFLDLVRTPQMRKCTLILMFAWFTSAVVYQGLVMRLGNLYIDFFISGVVELPGALLILLTIERLGRRLPFAASNIVAGVACLVTAFLPEGIAWLRTTVATLGRLGITMAFEIVYLVNSELYPTTLRNFGVSLCSGLCDFGGIIAPFLLFRLAAVWLELPLIIFGILASICGGLVMLLPETKGIALPETVDDVEK Hydrogen bonds contact Hydrophobic contact | ||||
| 70 | Dopamine beta-hydroxylase | 4ZEL | 6.47 | |
Target general information Gen name DBH Organism Homo sapiens (Human) Uniprot ID TTD ID NA Synonyms NA Protein family Copper type II ascorbate-dependent monooxygenase family Biochemical class Oxidoreductase Function Catalytic activity.Copper ion binding.Dopamine beta-monooxygenase activity.L-ascorbic acid binding. Related diseases Orthostatic hypotension 1 (ORTHYP1) [MIM:223360]: A form of orthostatic hypotension due to congenital dopamine beta-hydroxylase deficiency. Orthostatic hypotension, also known as postural hypotension, is a finding defined as a 20-mm Hg decrease in systolic pressure or a 10-mm Hg decrease in diastolic pressure occurring 3 minutes after a person has risen from supine to standing. Symptoms include dizziness, blurred vision, and sometimes syncope. ORTHYP1 is an autosomal recessive condition apparent from infancy or early childhood and characterized by low plasma and urinary levels of norepinephrine and epinephrine, and episodic hypoglycemia. {ECO:0000269|PubMed:11857564}. The disease is caused by variants affecting the gene represented in this entry. Drugs (DrugBank ID) DB00126; DB06774; DB09130; DB05394; DB00822; DB00988; DB00968; DB00550 Interacts with P00352; P63010-2; Q04656; Q8WUW1; Q9UNS2; Q71DI3; P61978; Q9Y2M5; Q92876; P08727; Q14693; P0DPK4; Q6GQQ9-2; P27986-2; Q9ULX5; Q96D59; Q8N6K7-2; Q9GZS3; Q8IUW3; Q86WT6-2 EC number 1.14.17.1 Uniprot keywords 3D-structure; Catecholamine biosynthesis; Copper; Cytoplasmic vesicle; Direct protein sequencing; Disease variant; Disulfide bond; Glycoprotein; Membrane; Metal-binding; Monooxygenase; Oxidoreductase; Proteomics identification; Reference proteome; Secreted; Signal-anchor; Transmembrane; Transmembrane helix; Vitamin C Protein physicochemical properties Chain ID A,B Molecular weight (Da) 123694 Length 1094 Aromaticity 0.1 Instability index 51.85 Isoelectric point 5.84 Charge (pH=7) -24.5 3D Binding mode Sequence PLPYHIPLDPEGSLELSWNVSYTQEAIHFQLLVRRLKAGVLFGMSDRGELENADLVVLWTDGDAYFADAWSDQKGQIHLDPQQDYQLLQVQRTPEGLTLLFKRPFGTCDPKDYLIEDGTVHLVYGILEEPFRSLEAINGSGLQMGLQRVQLLKPNIPEPELPSDACTMEVQAPNIQIPSQETTYWCYIKELPKGFSRHHIIKYEPIVTKGNEALVHHMEVFQCAPEMDSVPHFSGPCDSKMKPDRLNYCRHVLAAWALGAKAFYYPEEAGLAFGGPGSSRYLRLEVHYHNPLVIEGRNDSSGIRLYYTAKLRRFNAGIMELGLVYTPVMAIPPRETAFILTGYCTDKCTQLALPPSGIHIFASQLHTHLTGRKVVTVLVRDGREWEIVNQDNHYSPHFQEIRMLKKVVSVHPGDVLITSCTYNTEDRELATVGGFGILEEMCVNYVHYYPQTQLELCKSAVDAGFLQKYFHLINRFNNEDVCTCPQASVSQQFTSVPWNSFNRDVLKALYSFAPISMHCNKSSAVRFQGEWNLQPLPKVISTLEEPTVVSPLPYHIPLDPEGSLELSWNVSYTQEAIHFQLLVRRLKAGVLFGMSDRGELENADLVVLAYFADAWSDQKGQIHLDPQQDYQLLQVQRTPEGLTLLFKRPFGTCDPKDYLIEDGTVHLVYGILEEPFRSLEAINGSGLQMGLQRVQLLKPNIPEPELPSDACTMEVQAPNIQIPSQETTYWCYIKELPKGFSRHHIIKYEPIVTKGNEALVHHMEVFQCAPEVPHFSGPCDSKMLNYCRHVLAAWALGAKAFYYPEEAGLAFGGPGSSRYLRLEVHYHNPLVIEGRNDSSGIRLYYTAKLRRFNAGIMELGLVYTPVMAIPPRETAFILTGYCTDKCTQLALPPSGIHIFASQLHTHLTGRKVVTVLVRDGREWEIVNQDNHYSPHFQEIRMLKKVVSVHPGDVLITSCTYNTEDRELATVGGFGILEEMCVNYVHYYPQTQLELCKSAVDAGFLQKYFHLINRFNNEDVCTCPQASVSQQFTSVPWNSFNRDVLKALYSFAPISMHCNKSSAVRFQGEWNLQPLPKVISTLEEPTPQCVVSIGG Hydrogen bonds contact Hydrophobic contact | ||||
| 71 | SET domain containing 8 (KMT5A) | 5TEG | 6.46 | |
Target general information Gen name KMT5A Organism Homo sapiens (Human) Uniprot ID TTD ID Synonyms SETD8; SET8; SET07; SET domain-containing protein 8; PRSET7; PR/SET07; PR/SET domain-containing protein 07; PR-Set7; N-lysine methyltransferase KMT5A; Lysine-specific methylase 5A; Lysine N-methyltran Protein family Class V-like SAM-binding methyltransferase superfamily, Histone-lysine methyltransferase family, PR/SET subfamily Biochemical class Methyltransferase Function Specifically monomethylates 'Lys-20' of histone H4 (H4K20me1). H4K20me1 is enriched during mitosis and represents a specific tag for epigenetic transcriptional repression. Mainly functions in euchromatin regions, thereby playing a central role in the silencing of euchromatic genes. Required for cell proliferation, probably by contributing to the maintenance of proper higher-order structure of DNA during mitosis. Involved in chromosome condensation and proper cytokinesis. Nucleosomes are preferred as substrate compared to free histones. Mediates monomethylation of p53/TP53 at 'Lys-382', leading to repress p53/TP53-target genes. Plays a negative role in TGF-beta response regulation and a positive role in cell migration. Protein-lysine N-methyltransferase that monomethylates both histones and non-histone proteins. Related diseases Sick sinus syndrome 2 (SSS2) [MIM:163800]: The term 'sick sinus syndrome' encompasses a variety of conditions caused by sinus node dysfunction. The most common clinical manifestations are syncope, presyncope, dizziness, and fatigue. Electrocardiogram typically shows sinus bradycardia, sinus arrest, and/or sinoatrial block. Episodes of atrial tachycardias coexisting with sinus bradycardia ('tachycardia-bradycardia syndrome') are also common in this disorder. SSS occurs most often in the elderly associated with underlying heart disease or previous cardiac surgery, but can also occur in the fetus, infant, or child without heart disease or other contributing factors. SSS2 onset is in utero or at birth. {ECO:0000269|PubMed:15123648, ECO:0000269|PubMed:16407510, ECO:0000269|PubMed:20662977, ECO:0000269|PubMed:23103389}. The disease is caused by variants affecting the gene represented in this entry.; DISEASE: Brugada syndrome 8 (BRGDA8) [MIM:613123]: A tachyarrhythmia characterized by right bundle branch block and ST segment elevation on an electrocardiogram (ECG). It can cause the ventricles to beat so fast that the blood is prevented from circulating efficiently in the body. When this situation occurs, the individual will faint and may die in a few minutes if the heart is not reset. {ECO:0000269|PubMed:19165230}. The gene represented in this entry may be involved in disease pathogenesis.; DISEASE: Epilepsy, idiopathic generalized 18 (EIG18) [MIM:619521]: An autosomal dominant form of idiopathic generalized epilepsy, a disorder characterized by recurring generalized seizures in the absence of detectable brain lesions and/or metabolic abnormalities. Generalized seizures arise diffusely and simultaneously from both hemispheres of the brain. Seizure types include juvenile myoclonic seizures, absence seizures, and generalized tonic-clonic seizures. EIG18 is characterized by onset of myoclonic seizures in infancy. Although the seizures remit, some patients may have later speech or cognitive impairment. {ECO:0000269|PubMed:30127718}. Disease susceptibility is associated with variants affecting the gene represented in this entry. Drugs (DrugBank ID) NA Interacts with P62805; P07910; Q15672 EC number EC 2.1.1.- Uniprot keywords 3D-structure; Acetylation; Alternative splicing; Cell cycle; Cell division; Chromatin regulator; Chromosome; Coiled coil; Direct protein sequencing; Methyltransferase; Mitosis; Nucleus; Phosphoprotein; Proteomics identification; Reference proteome; Repressor; S-adenosyl-L-methionine; Transcription; Transcription regulation; Transferase; Ubl conjugation Protein physicochemical properties Chain ID A,D Molecular weight (Da) 19129.4 Length 167 Aromaticity 0.08 Instability index 49.18 Isoelectric point 7.88 Charge (pH=7) 1.37 3D Binding mode Sequence KSKAELQSEERKRIDELIESGKEEGMKIDLIDGKGRGVIATKQFSRGDFVVEYHGDLIEITDAKKREALYAQDPSTGCYMYYFQYLSKTYCVDATRETNRLGRLINHSKSGNCQTKLHDIDGVPHLILIASRDIAAGEELLYDYGDRSKASIEAHPWLKHKRHRVLR Hydrogen bonds contact Hydrophobic contact | ||||
| 72 | PRKR-like endoplasmic reticulum kinase (PERK) | 4G31 | 6.45 | |
Target general information Gen name EIF2AK3 Organism Homo sapiens (Human) Uniprot ID TTD ID Synonyms PEK Protein family Protein kinase superfamily, Ser/Thr protein kinase family, GCN2 subfamily Biochemical class Kinase Function Converts phosphorylated eIF-2-alpha/EIF2S1 either in a global protein synthesis inhibitor, leading to a reduced overall utilization of amino acids, or to a translation initiation activator of specific mRNAs, such as the transcriptional activator ATF4, and hence allowing ATF4-mediated reprogramming of amino acid biosynthetic gene expression to alleviate nutrient depletion. Serves as a critical effector of unfolded protein response (UPR)-induced G1 growth arrest due to the loss of cyclin-D1 (CCND1). Involved in control of mitochondrial morphology and function. Metabolic-stress sensing protein kinase that phosphorylates the alpha subunit of eukaryotic translation initiation factor 2 (eIF-2-alpha/EIF2S1) on 'Ser-52' during the unfolded protein response (UPR) and in response to low amino acid availability. Related diseases Wolcott-Rallison syndrome (WRS) [MIM:226980]: A rare autosomal recessive disorder, characterized by permanent neonatal or early infancy insulin-dependent diabetes and, at a later age, epiphyseal dysplasia, osteoporosis, growth retardation and other multisystem manifestations, such as hepatic and renal dysfunctions, intellectual disability and cardiovascular abnormalities. {ECO:0000269|PubMed:10932183, ECO:0000269|PubMed:12086964, ECO:0000269|PubMed:12960215, ECO:0000269|PubMed:16813601, ECO:0000269|PubMed:24168455, ECO:0000269|PubMed:24194294, ECO:0000269|PubMed:27145240, ECO:0000269|PubMed:28220546, ECO:0000269|PubMed:30906465, ECO:0000269|PubMed:30922274, ECO:0000269|PubMed:32216767, ECO:0000269|PubMed:34123975}. The disease is caused by variants affecting the gene represented in this entry. Drugs (DrugBank ID) NA Interacts with Q9NZJ5; P11021 EC number EC 2.7.11.1 Uniprot keywords 3D-structure; ADP-ribosylation; ATP-binding; Diabetes mellitus; Disease variant; Endoplasmic reticulum; Glycoprotein; Kinase; Membrane; Nucleotide-binding; Phosphoprotein; Proteomics identification; Reference proteome; Serine/threonine-protein kinase; Signal; Stress response; Transferase; Translation regulation; Transmembrane; Transmembrane helix; Tyrosine-protein kinase; Unfolded protein response Protein physicochemical properties Chain ID A Molecular weight (Da) 29033.5 Length 248 Aromaticity 0.1 Instability index 43.71 Isoelectric point 7.75 Charge (pH=7) 1.27 3D Binding mode Sequence GRYLTDFEPIQCLGRGGVVFEAKNKVDDCNYAIKRIRLPNRELAREKVMREVKALAKLEHPGIVRYFNAWLEKNKVYLYIQMQLCRKENLKDWMNGRCTIEERERSVCLHIFLQIAEAVEFLHSKGLMHRDLKPSNIFFTMDDVVKVGDFGLVGTKLYMSPEQIHGNSYSHKVDIFSLGLILFELLYPFSTQMERVRTLTDVRNLKFPPLFTQKYPCEYVMVQDMLSPSPMERPEAINIIENAVFEDL Hydrogen bonds contact Hydrophobic contact | ||||
| 73 | Pyruvate kinase PKLR | 4IP7 | 6.45 | |
Target general information Gen name PKLR Organism Homo sapiens (Human) Uniprot ID TTD ID NA Synonyms PKL;PK1 Protein family Pyruvate kinase family Biochemical class Transferase Function ATP binding.Kinase activity.Magnesium ion binding.Potassium ion binding.Pyruvate kinase activity. Related diseases Pyruvate kinase hyperactivity (PKHYP) [MIM:102900]: Autosomal dominant phenotype characterized by increase of red blood cell ATP. {ECO:0000269|PubMed:9090535}. The disease is caused by variants affecting the gene represented in this entry.; DISEASE: Pyruvate kinase deficiency of red cells (PKRD) [MIM:266200]: A frequent cause of hereditary non-spherocytic hemolytic anemia. Clinically, pyruvate kinase-deficient patients suffer from a highly variable degree of chronic hemolysis, ranging from severe neonatal jaundice and fatal anemia at birth, severe transfusion-dependent chronic hemolysis, moderate hemolysis with exacerbation during infection, to a fully compensated hemolysis without apparent anemia. {ECO:0000269|PubMed:10087985, ECO:0000269|PubMed:10772876, ECO:0000269|PubMed:11328279, ECO:0000269|PubMed:11960989, ECO:0000269|PubMed:1536957, ECO:0000269|PubMed:1896471, ECO:0000269|PubMed:19085939, ECO:0000269|PubMed:2018831, ECO:0000269|PubMed:21794208, ECO:0000269|PubMed:7706479, ECO:0000269|PubMed:8161798, ECO:0000269|PubMed:8180378, ECO:0000269|PubMed:8476433, ECO:0000269|PubMed:8481523, ECO:0000269|PubMed:8483951, ECO:0000269|PubMed:8664896, ECO:0000269|PubMed:8807089, ECO:0000269|PubMed:9075576, ECO:0000269|PubMed:9482576, ECO:0000269|PubMed:9827908, ECO:0000269|PubMed:9886305, ECO:0000269|Ref.24}. The disease is caused by variants affecting the gene represented in this entry. Drugs (DrugBank ID) DB02726; DB00787; DB04551; DB16236; DB00119 Interacts with Q9UBL6-2 EC number 2.7.1.40 Uniprot keywords 3D-structure; Allosteric enzyme; Alternative splicing; ATP-binding; Disease variant; Glycolysis; Hereditary hemolytic anemia; Kinase; Magnesium; Manganese; Metal-binding; Nucleotide-binding; Phosphoprotein; Potassium; Proteomics identification; Pyruvate; Reference proteome; Transferase Protein physicochemical properties Chain ID A,B,C,D Molecular weight (Da) 45695.1 Length 421 Aromaticity 0.06 Instability index 34.44 Isoelectric point 6.88 Charge (pH=7) -0.35 3D Binding mode Sequence GTAFFQQQQLPAAMADTFLEHLCLLDIDSEPVAARSTSIIATIGPASRSVERLKEMIKAGMNIARLNFSHGSHEYHAESIANVREAVESFSPLSYRPVAIALDTKGPEIGLSEQDVRDLRFGVEHGVDIVFASFVRKASDVAAVRAALGPEGHGIKIISKIENHEGVKRFDEILEVSDGIMVARGDLGIEIPAEKVFLAQKMMIGRCNLAGKPVVCATQMLESMITKPRPTRAETSDVANAVLDGADCIMLSGETAKGNFPVEAVKMQHAIAREAEAAVYHRQLFEELRRAAPLSRDPTEVTAIGAVEAAFKCCAAAIIVLTTTGRSAQLLSRYRPRAAVIAVTRSAQAARQVHLCRGVFPLLYREPPEAIWADDVDRRVQFGIESGKLRGFLRVGDLVIVVTGWRPGSGYTNIMRVLSIS Hydrogen bonds contact Hydrophobic contact | ||||
| 74 | Acyloxyacyl hydrolase (neutrophil) | 5W7C | 6.45 | |
Target general information Gen name AOAH Organism Homo sapiens (Human) Uniprot ID TTD ID Synonyms Acyloxyacyl hydrolase Protein family NA Biochemical class NA Function Removes the secondary (acyloxyacyl-linked) fatty acyl chains from the lipid A region of bacterial lipopolysaccharides. By breaking down LPS, terminates the host response to bacterial infection and prevents prolonged and damaging inflammatory responses (By similarity). In peritoneal macrophages, seems to be important for recovery from a state of immune tolerance following infection by Gram-negative bacteria (By similarity). Related diseases Major depressive disorder (MDD) [MIM:608516]: A common psychiatric disorder. It is a complex trait characterized by one or more major depressive episodes without a history of manic, mixed, or hypomanic episodes. A major depressive episode is characterized by at least 2 weeks during which there is a new onset or clear worsening of either depressed mood or loss of interest or pleasure in nearly all activities. Four additional symptoms must also be present including changes in appetite, weight, sleep, and psychomotor activity; decreased energy; feelings of worthlessness or guilt; difficulty thinking, concentrating, or making decisions; or recurrent thoughts of death or suicidal ideation, plans, or attempts. The episode must be accompanied by distress or impairment in social, occupational, or other important areas of functioning. {ECO:0000269|PubMed:15229186}. Disease susceptibility is associated with variants affecting the gene represented in this entry. Drugs (DrugBank ID) NA Interacts with Q15700 EC number EC 3.1.1.77 Uniprot keywords 3D-structure; Alternative splicing; Calcium; Cytoplasmic vesicle; Direct protein sequencing; Disulfide bond; Glycoprotein; Hydrolase; Lipid metabolism; Metal-binding; Proteomics identification; Reference proteome; Secreted; Signal; Zymogen Protein physicochemical properties Chain ID C Molecular weight (Da) 47779.7 Length 420 Aromaticity 0.1 Instability index 43.45 Isoelectric point 7.72 Charge (pH=7) 2.1 3D Binding mode Sequence GSDICSLPVLAKICQKIKLAMEQSVPFKDVDSDKYSVFPTLRGYHWRGRDCNDSDESVYPGRRPNNWDVHQDSNCNGIWGVDPKDGVPYEKKFCEGSQPRGIILLGDAAGAHFHISPEWITASQMSLNSFINLPTALTNELDWPQLSGATGFLDSTVGIKEKSIYLRLWKRNHCNHRDYQNISRNGASSRNLKKFIESLSRNKVLDYPAIVIYAMIGNDVCSGKSDPVPAMTTPEKLYSNVMQTLKHLNSHLPNGSHVILYGLPDGTFLWDNLHNRYHPLGQLNKDMTYAQLYSFLNCLQVSPCHGWMSSNKTLRTLTSERAEQLSNTLKKIAASEKFTNFNLFYMDFAFHEIIQEWQKRGGQPWQLIEPVDGFHPNEVALLLLADHFWKKVQLQWPQILGKENPFNPQIKQVFGDQGGH Hydrogen bonds contact Hydrophobic contact | ||||
| 75 | Bifunctional dihydrofolate reductase-thymidylate synthase | 1J3K | 6.45 | |
Target general information Gen name N/A Organism Plasmodium falciparum (isolate K1 / Thailand) Uniprot ID TTD ID NA Synonyms NA Protein family Dihydrofolate reductase family; Thymidylate synthase family Biochemical class Oxidoreductase Function Dihydrofolate reductase activity.Thymidylate synthase activity. Related diseases Acute hepatic porphyria (AHEPP) [MIM:612740]: A form of porphyria. Porphyrias are inherited defects in the biosynthesis of heme, resulting in the accumulation and increased excretion of porphyrins or porphyrin precursors. They are classified as erythropoietic or hepatic, depending on whether the enzyme deficiency occurs in red blood cells or in the liver. AHP is characterized by attacks of gastrointestinal disturbances, abdominal colic, paralyses and peripheral neuropathy. Most attacks are precipitated by drugs, alcohol, caloric deprivation, infections, or endocrine factors. {ECO:0000269|PubMed:10706561, ECO:0000269|PubMed:1309003, ECO:0000269|PubMed:1569184, ECO:0000269|PubMed:17236137, ECO:0000269|PubMed:2063868}. The disease is caused by variants affecting the gene represented in this entry. Drugs (DrugBank ID) DB01131; DB00205; DB01299 Interacts with NA EC number 1.5.1.3; 2.1.1.45 Uniprot keywords 3D-structure; Methyltransferase; Multifunctional enzyme; NADP; Nucleotide biosynthesis; One-carbon metabolism; Oxidoreductase; Transferase Protein physicochemical properties Chain ID A,B Molecular weight (Da) 61720.6 Length 525 Aromaticity 0.13 Instability index 33.9 Isoelectric point 8.79 Charge (pH=7) 11.61 3D Binding mode Sequence NSIHPNDFQIYNSLKYKYHPEYQYLNIIYDIMMNGNKQSDRTGVGVLSKFGYIMKFDLSQYFPLLTTKKLFLRGIIEELLWFIRGETNGNTLLNKNVRIWEANGTREFLDNRKLFHREVNDLGPIYGFQWRHFGAEYTNMYDNYENKGVDQLKNIINLIKNDPTSRRILLCAWNVKDLDQMALPPCHILCQFYVFDGKLSCIMYQRSCDLGLGVPFNIASYSIFTHMIAQVCNLQPAQFIHVLGNAHVYNNHIDSLKIQLNRIPYPFPTLKLNPDIKNIEDFTISDFTIQNYVHHEKISMDMAAMMEQVCDVFDIYAICACCKVESKNEGKKNEVFNNYTFRGLGNKGVLPWKCISLDMKYFRAVTTYVNESKYEKLKYKRCKYLPNSKKLQNVVVMGRTNWESIPKKFKPLSNRINVILSRTLKKEDFDEDVYIINKVEDLIVLLGKLNYYKCFILGGSVVYQEFLEKKLIKKIYFTRINSTYECDVFFPEINENEYQIISVSDVYTSNNTTLDFIIYKKTNNK Hydrogen bonds contact Hydrophobic contact | ||||
| 76 | Vascular endothelial growth factor receptor 1 (FLT-1) | 5EX3 | 6.45 | |
Target general information Gen name FLT1 Organism Homo sapiens (Human) Uniprot ID TTD ID Synonyms Vascular permeability factor receptor; VEGFR1; VEGFR-1; VEGF-1 receptor; Tyrosine-protein kinase receptor FLT; Tyrosine-protein kinase FRT; Fms-like tyrosine kinase 1; FRT; FLT Protein family Protein kinase superfamily, Tyr protein kinase family, CSF-1/PDGF receptor subfamily Biochemical class Kinase Function May play an essential role as a negative regulator of embryonic angiogenesis by inhibiting excessive proliferation of endothelial cells. Can promote endothelial cell proliferation, survival and angiogenesis in adulthood. Its function in promoting cell proliferation seems to be cell-type specific. Promotes PGF-mediated proliferation of endothelial cells, proliferation of some types of cancer cells, but does not promote proliferation of normal fibroblasts (in vitro). Has very high affinity for VEGFA and relatively low protein kinase activity; may function as a negative regulator of VEGFA signaling by limiting the amount of free VEGFA and preventing its binding to KDR. Likewise, isoforms lacking a transmembrane domain, such as isoform 2, isoform 3 and isoform 4, may function as decoy receptors for VEGFA. Modulates KDR signaling by forming heterodimers with KDR. Ligand binding leads to the activation of several signaling cascades. Activation of PLCG leads to the production of the cellular signaling molecules diacylglycerol and inositol 1,4,5-trisphosphate and the activation of protein kinase C. Mediates phosphorylation of PIK3R1, the regulatory subunit of phosphatidylinositol 3-kinase, leading to activation of phosphatidylinositol kinase and the downstream signaling pathway. Mediates activation of MAPK1/ERK2, MAPK3/ERK1 and the MAP kinase signaling pathway, as well as of the AKT1 signaling pathway. Phosphorylates SRC and YES1, and may also phosphorylate CBL. Isoform 1 phosphorylates PLCG. Promotes phosphorylation of AKT1 at 'Ser-473'. Promotes phosphorylation of PTK2/FAK1. Isoform 7 has a truncated kinase domain; it increases phosphorylation of SRC at 'Tyr-418' by unknown means and promotes tumor cell invasion. Tyrosine-protein kinase that acts as a cell-surface receptor for VEGFA, VEGFB and PGF, and plays an essential role in the development of embryonic vasculature, the regulation of angiogenesis, cell survival, cell migration, macrophage function, chemotaxis, and cancer cell invasion. Related diseases Can contribute to cancer cell survival, proliferation, migration, and invasion, and tumor angiogenesis and metastasis. May contribute to cancer pathogenesis by promoting inflammatory responses and recruitment of tumor-infiltrating macrophages.; DISEASE: Abnormally high expression of soluble isoforms (isoform 2, isoform 3 or isoform 4) may be a cause of preeclampsia. Drugs (DrugBank ID) DB06626; DB05932; DB10770; DB12010; DB11679; DB06101; DB09078; DB06080; DB07288; DB09079; DB05913; DB06589; DB09221; DB08896; DB15685; DB00398; DB01268; DB05075; DB11800; DB04879 Interacts with P22681; P46109; P49763; P27986; Q12913; P15692; P15692-4; PRO_0000391621 [P98160]; PRO_0000391622 [P98160] EC number EC 2.7.10.1 Uniprot keywords 3D-structure; Alternative splicing; Angiogenesis; ATP-binding; Cell membrane; Chemotaxis; Cytoplasm; Developmental protein; Differentiation; Direct protein sequencing; Disulfide bond; Endosome; Glycoprotein; Immunoglobulin domain; Kinase; Membrane; Nucleotide-binding; Phosphoprotein; Proteomics identification; Receptor; Reference proteome; Repeat; Secreted; Signal; Transferase; Transmembrane; Transmembrane helix; Tyrosine-protein kinase; Ubl conjugation Protein physicochemical properties Chain ID D Molecular weight (Da) 49666.3 Length 435 Aromaticity 0.08 Instability index 43.84 Isoelectric point 7.05 Charge (pH=7) 0.14 3D Binding mode Sequence MEPLKVEKFATANRGNGLRAVTPLRPGELLFRSDPLAYTVCKGSRGVVCDRCLLGKEKLMRCSQCRVAKYCSAKCQKKAWPDHKRECKCLKSCKPRYPPDSVRLLGRVVFKLMDGAPSESEKLYSFYDLESNINKLTEDRKEGLRQLVMTFQHFMREEIQDASQLPPAFDLFEAFAKVICNSFTICNAEMQEVGVGLYPSISLLNHSCDPNCSIVFNGPHLLLRAVRDIEVGEELTICYLDMLMTSEERRKQLRDQYCFECDCFRCQTQDKDADMLTGDEQVWKEVQESLKKIEELKAHWKWEQVLAMCQAIISSNSERLPDINIYQLKVLDCAMDACINLGLLEEALFYGTRTMEPYRIFFPGSHPVRGVQVMKVGKLQLHQGMFPQAMKNLRLAFDIMRVTHGREHSLIEDLILLLEECDANIRASKLGXSLG Hydrogen bonds contact Hydrophobic contact | ||||
| 77 | Alpha-L-iduronidase (IDUA) | 6I6R | 6.44 | |
Target general information Gen name IDUA Organism Homo sapiens (Human) Uniprot ID TTD ID Synonyms NA Protein family Glycosyl hydrolase 39 family Biochemical class Glycosyl hydrolase Function extracellular exosome, lysosomal lumen, hydrolase activity, hydrolyzing O-glycosyl compounds, L-iduronidase activity, chondroitin sulfate catabolic process, dermatan sulfate catabolic process, disaccharide metabolic process, glycosaminoglycan catabolic process, heparin catabolic process Related diseases Mucopolysaccharidosis 1H (MPS1H) [MIM:607014]: A severe form of mucopolysaccharidosis type 1, a rare lysosomal storage disease characterized by progressive physical deterioration with urinary excretion of dermatan sulfate and heparan sulfate. Patients with MPS1H usually present, within the first year of life, a combination of hepatosplenomegaly, skeletal deformities, corneal clouding and severe intellectual disability. Obstructive airways disease, respiratory infection and cardiac complications usually result in death before 10 years of age. {ECO:0000269|PubMed:10466419, ECO:0000269|PubMed:10735634, ECO:0000269|PubMed:12559846, ECO:0000269|PubMed:1301941, ECO:0000269|PubMed:15300847, ECO:0000269|PubMed:19396826, ECO:0000269|PubMed:21394825, ECO:0000269|PubMed:24036510, ECO:0000269|PubMed:31194252, ECO:0000269|PubMed:7550232, ECO:0000269|PubMed:7550242, ECO:0000269|PubMed:7951228, ECO:0000269|PubMed:8019563, ECO:0000269|PubMed:8328452, ECO:0000269|PubMed:8401515, ECO:0000269|Ref.20}. The disease is caused by variants affecting the gene represented in this entry.; DISEASE: Mucopolysaccharidosis 1H/S (MPS1H/S) [MIM:607015]: A form of mucopolysaccharidosis type 1, a rare lysosomal storage disease characterized by progressive physical deterioration with urinary excretion of dermatan sulfate and heparan sulfate. MPS1H/S represents an intermediate phenotype of the MPS1 clinical spectrum. It is characterized by relatively little neurological involvement, but most of the somatic symptoms described for severe MPS1 develop in the early to mid-teens, causing considerable loss of mobility. {ECO:0000269|PubMed:10466419, ECO:0000269|PubMed:10735634, ECO:0000269|PubMed:12559846, ECO:0000269|PubMed:15300847, ECO:0000269|PubMed:21394825, ECO:0000269|PubMed:7550232, ECO:0000269|PubMed:7550242, ECO:0000269|PubMed:8401515}. The disease is caused by variants affecting the gene represented in this entry.; DISEASE: Mucopolysaccharidosis 1S (MPS1S) [MIM:607016]: A mild form of mucopolysaccharidosis type 1, a rare lysosomal storage disease characterized by progressive physical deterioration with urinary excretion of dermatan sulfate and heparan sulfate. Patients with MPS1S may have little or no neurological involvement, normal stature and life span, but present development of joints stiffness, mild hepatosplenomegaly, aortic valve disease and corneal clouding. {ECO:0000269|PubMed:12559846, ECO:0000269|PubMed:15300847, ECO:0000269|PubMed:19396826, ECO:0000269|PubMed:21394825, ECO:0000269|PubMed:25256405, ECO:0000269|PubMed:7550232, ECO:0000269|PubMed:7550242, ECO:0000269|PubMed:8213840}. The disease is caused by variants affecting the gene represented in this entry. Drugs (DrugBank ID) DB09301 Interacts with NA EC number EC 3.2.1.76 Uniprot keywords 3D-structure; Alternative splicing; Disease variant; Disulfide bond; Glycoprotein; Glycosidase; Hydrolase; Lysosome; Mucopolysaccharidosis; Proteomics identification; Reference proteome; Signal Protein physicochemical properties Chain ID A Molecular weight (Da) 67045.6 Length 597 Aromaticity 0.11 Instability index 38.21 Isoelectric point 8.9 Charge (pH=7) 7.67 3D Binding mode Sequence EAPHLVQVDAARALWPLRRFWRSTGFCPPLYVLSWDQQLNLAYVGAVPHRGIKQVRTHWLLELVTTRGLSYNFTHLDGYLDLLRENQLLPGFELMGSASGHFTDFEDKQQVFEWKDLVSSLARRYIGRYGLAHVSKWNFETWNEPDHHDFDNVSMTMQGFLNYYDACSEGLRAASPALRLGGPGDSFHTPPRSPLSWGLLRHCHDGTNFFTGEAGVRLDYISLHRKGARSSISILEQEKVVAQQIRQLFPKFADTPIYNDEADPLVGWSLPQPWRADVTYAAMVVKVIAQHQNLLLAAFPYALLSNDNAFLSYHPHPFAQRTLTARFQVNNTRPPHVQLLRKPVLTAMGLLALLDEEQLWAEVSQAGTVLDSNHTVGVLASAHRPQGPADAWRAAVLIYASDDTRAHPNRSVAVTLRLRGVPPGPGLVYVTRYLDNGLCSPDGEWRRLGRPVFPTAEQFRRMRAAEDPVAAAPRPLPAGGRLTLRPALRLPSLLLVHVCARPEKPPGQVTRLRALPLTQGQLVLVWSDEHVGSKCLWTYEIQFSQAYTPVSRKPSTFNLFVFSPDTGAVSGSYRVRALDYWARPGPFSDPVPYLEVP Hydrogen bonds contact Hydrophobic contact | ||||
| 78 | Platelet-derived growth factor receptor alpha (PDGFRA) | 5K5X | 6.44 | |
Target general information Gen name PDGFRA Organism Homo sapiens (Human) Uniprot ID TTD ID Synonyms RHEPDGFRA; Platelet-derived growth factor receptor 2; Platelet-derived growth factor alpha receptor; PDGFR2; PDGFR-alpha; PDGFR-2; PDGF-R-alpha; CD140a antigen; CD140a; CD140 antigen-like family membe Protein family Protein kinase superfamily, Tyr protein kinase family, CSF-1/PDGF receptor subfamily Biochemical class Kinase Function Depending on the context, promotes or inhibits cell proliferation and cell migration. Plays an important role in the differentiation of bone marrow-derived mesenchymal stem cells. Required for normal skeleton development and cephalic closure during embryonic development. Required for normal development of the mucosa lining the gastrointestinal tract, and for recruitment of mesenchymal cells and normal development of intestinal villi. Plays a role in cell migration and chemotaxis in wound healing. Plays a role in platelet activation, secretion of agonists from platelet granules, and in thrombin-induced platelet aggregation. Binding of its cognate ligands - homodimeric PDGFA, homodimeric PDGFB, heterodimers formed by PDGFA and PDGFB or homodimeric PDGFC -leads to the activation of several signaling cascades; the response depends on the nature of the bound ligand and is modulated by the formation of heterodimers between PDGFRA and PDGFRB. Phosphorylates PIK3R1, PLCG1, and PTPN11. Activation of PLCG1 leads to the production of the cellular signaling molecules diacylglycerol and inositol 1,4,5-trisphosphate, mobilization of cytosolic Ca(2+) and the activation of protein kinase C. Phosphorylates PIK3R1, the regulatory subunit of phosphatidylinositol 3-kinase, and thereby mediates activation of the AKT1 signaling pathway. Mediates activation of HRAS and of the MAP kinases MAPK1/ERK2 and/or MAPK3/ERK1. Promotes activation of STAT family members STAT1, STAT3 and STAT5A and/or STAT5B. Receptor signaling is down-regulated by protein phosphatases that dephosphorylate the receptor and its down-stream effectors, and by rapid internalization of the activated receptor. Tyrosine-protein kinase that acts as a cell-surface receptor for PDGFA, PDGFB and PDGFC and plays an essential role in the regulation of embryonic development, cell proliferation, survival and chemotaxis. Related diseases A chromosomal aberration involving PDGFRA is found in some cases of hypereosinophilic syndrome. Interstitial chromosomal deletion del(4)(q12q12) causes the fusion of FIP1L1 and PDGFRA (FIP1L1-PDGFRA). Mutations that cause overexpression and/or constitutive activation of PDGFRA may be a cause of hypereosinophilic syndrome. {ECO:0000269|PubMed:12808148}.; DISEASE: Gastrointestinal stromal tumor (GIST) [MIM:606764]: Common mesenchymal neoplasms arising in the gastrointestinal tract, most often in the stomach. They are histologically, immunohistochemically, and genetically different from typical leiomyomas, leiomyosarcomas, and schwannomas. Most GISTs are composed of a fairly uniform population of spindle-shaped cells. Some tumors are dominated by epithelioid cells or contain a mixture of spindle and epithelioid morphologies. Primary GISTs in the gastrointestinal tract commonly metastasize in the omentum and mesenteries, often as multiple nodules. However, primary tumors may also occur outside of the gastrointestinal tract, in other intra-abdominal locations, especially in the omentum and mesentery. {ECO:0000269|PubMed:12522257, ECO:0000269|PubMed:15928335}. The gene represented in this entry may be involved in disease pathogenesis. Mutations causing PDGFRA constitutive activation have been found in gastrointestinal stromal tumors lacking KIT mutations (PubMed:12522257). {ECO:0000269|PubMed:12522257}.; DISEASE: GIST-plus syndrome (GISTPS) [MIM:175510]: A disorder characterized by multiple mesenchymal tumors of the gastrointestinal tract, including gastrointestinal stromal tumor, inflammatory fibroid polyps, and fibroid tumors. Additional features are coarse facies and skin, broad hands and feet, and premature tooth loss. GISTPS is an autosomal dominant disease with incomplete penetrance. Gastrointestinal stromal tumor and inflammatory fibroid polyps may also occur in isolation. {ECO:0000269|PubMed:14699510, ECO:0000269|PubMed:17087943, ECO:0000269|PubMed:25975287}. The disease is caused by variants affecting the gene represented in this entry. Drugs (DrugBank ID) DB12742; DB00102; DB12147; DB10772; DB12010; DB00619; DB09078; DB06595; DB09079; DB06043; DB06589; DB08901; DB08896; DB14840; DB01268; DB11800; DB05146 Interacts with P46108; P46109; P00533; Q8N6L0; P04085; P01127; Q9NRA1; P31947; P62258; Q9NRA1-1; A8T7D5; P05067; Q8IY26 EC number EC 2.7.10.1 Uniprot keywords 3D-structure; Alternative splicing; ATP-binding; Cell membrane; Cell projection; Chemotaxis; Developmental protein; Disease variant; Disulfide bond; Glycoprotein; Golgi apparatus; Host-virus interaction; Immunoglobulin domain; Kinase; Membrane; Nucleotide-binding; Phosphoprotein; Proteomics identification; Proto-oncogene; Receptor; Reference proteome; Repeat; Signal; Transferase; Transmembrane; Transmembrane helix; Tyrosine-protein kinase; Ubl conjugation Protein physicochemical properties Chain ID A Molecular weight (Da) 39294.8 Length 345 Aromaticity 0.11 Instability index 46.74 Isoelectric point 6.6 Charge (pH=7) -1.38 3D Binding mode Sequence RYEIRWRVIESISPDGHEYIYVDPMQLPYDSRWEFPRDGLVLGRVLGSGAFGKVVEGTAYGLSRSQPVMKVAVKMLKPTARSSEKQALMSELKIMTHLGPHLNIVNLLGACTKSGPIYIITEYCFYGDLVNYLHKNRDSFLSHSMLDSEVKNLLSDDNSEGLTLLDLLSFTYQVARGMEFLASKNCVHRDLAARNVLLAQGKIVKICDFGLARDIMHDSNYVSKGSTFLPVKWMAPESIFDNLYTTLSDVWSYGILLWEIFSLGGTPYPGMMVDSTFYNKIKSGYRMAKPDHATSEVYEIMVKCWNSEPEKRPSFYHLSEIVENLLPGQYKKSYEKIHLDFLKSD Hydrogen bonds contact Hydrophobic contact | ||||
| 79 | Protein kinase C epsilon (PRKCE) | 5LIH | 6.44 | |
Target general information Gen name PRKCE Organism Homo sapiens (Human) Uniprot ID TTD ID Synonyms Protein kinase C epsilon type; PKCE; PKC epsilon; NPKC-epsilon Protein family Protein kinase superfamily, AGC Ser/Thr protein kinase family, PKC subfamily Biochemical class Kinase Function Mediates cell adhesion to the extracellular matrix via integrin-dependent signaling, by mediating angiotensin-2-induced activation of integrin beta-1 (ITGB1) in cardiac fibroblasts. Phosphorylates MARCKS, which phosphorylates and activates PTK2/FAK, leading to the spread of cardiomyocytes. Involved in the control of the directional transport of ITGB1 in mesenchymal cells by phosphorylating vimentin (VIM), an intermediate filament (IF) protein. In epithelial cells, associates with and phosphorylates keratin-8 (KRT8), which induces targeting of desmoplakin at desmosomes and regulates cell-cell contact. Phosphorylates IQGAP1, which binds to CDC42, mediating epithelial cell-cell detachment prior to migration. In HeLa cells, contributes to hepatocyte growth factor (HGF)-induced cell migration, and in human corneal epithelial cells, plays a critical role in wound healing after activation by HGF. During cytokinesis, forms a complex with YWHAB, which is crucial for daughter cell separation, and facilitates abscission by a mechanism which may implicate the regulation of RHOA. In cardiac myocytes, regulates myofilament function and excitation coupling at the Z-lines, where it is indirectly associated with F-actin via interaction with COPB1. During endothelin-induced cardiomyocyte hypertrophy, mediates activation of PTK2/FAK, which is critical for cardiomyocyte survival and regulation of sarcomere length. Plays a role in the pathogenesis of dilated cardiomyopathy via persistent phosphorylation of troponin I (TNNI3). Involved in nerve growth factor (NFG)-induced neurite outgrowth and neuron morphological change independently of its kinase activity, by inhibition of RHOA pathway, activation of CDC42 and cytoskeletal rearrangement. May be involved in presynaptic facilitation by mediating phorbol ester-induced synaptic potentiation. Phosphorylates gamma-aminobutyric acid receptor subunit gamma-2 (GABRG2), which reduces the response of GABA receptors to ethanol and benzodiazepines and may mediate acute tolerance to the intoxicating effects of ethanol. Upon PMA treatment, phosphorylates the capsaicin- and heat-activated cation channel TRPV1, which is required for bradykinin-induced sensitization of the heat response in nociceptive neurons. Is able to form a complex with PDLIM5 and N-type calcium channel, and may enhance channel activities and potentiates fast synaptic transmission by phosphorylating the pore-forming alpha subunit CACNA1B (CaV2. 2). In prostate cancer cells, interacts with and phosphorylates STAT3, which increases DNA-binding and transcriptional activity of STAT3 and seems to be essential for prostate cancer cell invasion. Downstream of TLR4, plays an important role in the lipopolysaccharide (LPS)-induced immune response by phosphorylating and activating TICAM2/TRAM, which in turn activates the transcription factor IRF3 and subsequent cytokines production. In differentiating erythroid progenitors, is regulated by EPO and controls the protection against the TNFSF10/TRAIL-mediated apoptosis, via BCL2. May be involved in the regulation of the insulin-induced phosphorylation and activation of AKT1. Calcium-independent, phospholipid- and diacylglycerol (DAG)-dependent serine/threonine-protein kinase that plays essential roles in the regulation of multiple cellular processes linked to cytoskeletal proteins, such as cell adhesion, motility, migration and cell cycle, functions in neuron growth and ion channel regulation, and is involved in immune response, cancer cell invasion and regulation of apoptosis. Related diseases Hyperinsulinemic hypoglycemia, familial, 2 (HHF2) [MIM:601820]: A form of hyperinsulinemic hypoglycemia, a clinically and genetically heterogeneous disorder characterized by inappropriate insulin secretion from the pancreatic beta-cells in the presence of low blood glucose levels. HHF2 is a common cause of persistent hypoglycemia in infancy. Unless early and aggressive intervention is undertaken, brain damage from recurrent episodes of hypoglycemia may occur. HHF2 inheritance can be autosomal dominant or autosomal recessive. {ECO:0000269|PubMed:10204114, ECO:0000269|PubMed:12364426, ECO:0000269|PubMed:15562009, ECO:0000269|PubMed:15579781, ECO:0000269|PubMed:15807877, ECO:0000269|PubMed:15998776, ECO:0000269|PubMed:16332676, ECO:0000269|PubMed:16357843, ECO:0000269|PubMed:18596924, ECO:0000269|PubMed:19357197, ECO:0000269|PubMed:7847376, ECO:0000269|PubMed:8923010}. The disease is caused by variants affecting the gene represented in this entry.; DISEASE: Diabetes mellitus, permanent neonatal, 2 (PNDM2) [MIM:618856]: A form of permanent neonatal diabetes mellitus, a type of diabetes characterized by onset of persistent hyperglycemia within the first six months of life. Initial clinical manifestations include intrauterine growth retardation, hyperglycemia, glycosuria, osmotic polyuria, severe dehydration, and failure to thrive. Some PNDM2 patients may also have developmental delay, muscle weakness, epilepsy and dysmorphic features. PNDM2 transmission pattern is consistent with autosomal dominant inheritance. {ECO:0000269|PubMed:15115830, ECO:0000269|PubMed:15292329, ECO:0000269|PubMed:15448106, ECO:0000269|PubMed:15448107, ECO:0000269|PubMed:15580558, ECO:0000269|PubMed:15583126, ECO:0000269|PubMed:16609879, ECO:0000269|PubMed:16731833, ECO:0000269|PubMed:17213273, ECO:0000269|PubMed:17652641, ECO:0000269|PubMed:17855752, ECO:0000269|PubMed:20022885, ECO:0000269|PubMed:28842488}. The disease is caused by variants affecting the gene represented in this entry.; DISEASE: Transient neonatal diabetes mellitus 3 (TNDM3) [MIM:610582]: Neonatal diabetes mellitus, defined as insulin-requiring hyperglycemia within the first month of life, is a rare entity. In about half of the neonates, diabetes is transient and resolves at a median age of 3 months, whereas the rest have a permanent form of diabetes. In a significant number of patients with transient neonatal diabetes mellitus, diabetes type 2 appears later in life. The onset and severity of TNDM3 is variable with childhood-onset diabetes, gestational diabetes or adult-onset diabetes described. {ECO:0000269|PubMed:15718250, ECO:0000269|PubMed:15784703}. The disease is caused by variants affecting the gene represented in this entry.; DISEASE: Defects in KCNJ11 may contribute to non-insulin-dependent diabetes mellitus (NIDDM), also known as diabetes mellitus type 2.; DISEASE: Maturity-onset diabetes of the young 13 (MODY13) [MIM:616329]: A form of diabetes that is characterized by an autosomal dominant mode of inheritance, onset in childhood or early adulthood (usually before 25 years of age), a primary defect in insulin secretion and frequent insulin-independence at the beginning of the disease. {ECO:0000269|PubMed:22701567}. The disease is caused by variants affecting the gene represented in this entry. Drugs (DrugBank ID) DB09096; DB11752; DB04209; DB12010; DB06064; DB02010; DB00675 Interacts with P05067; P15056; P08238; C6GKH1; Q9BPZ7; P17252; Q15349; P31947; P37840; P62258; P63104 EC number EC 2.7.11.13 Uniprot keywords 3D-structure; ATP-binding; Cell adhesion; Cell cycle; Cell division; Cell membrane; Cytoplasm; Cytoskeleton; Immunity; Kinase; Membrane; Metal-binding; Nucleotide-binding; Nucleus; Phosphoprotein; Proteomics identification; Reference proteome; Repeat; Serine/threonine-protein kinase; Transferase; Zinc; Zinc-finger Protein physicochemical properties Chain ID A,F Molecular weight (Da) 39129.4 Length 339 Aromaticity 0.12 Instability index 42.21 Isoelectric point 6.17 Charge (pH=7) -3.73 3D Binding mode Sequence SLGLQDFDLLRVIGRGSYAKVLLVRLKKTDRIYAMKVVKKELVWVQTEKHVFEQASNHPFLVGLHSCFQTESRLFFVIEYVNGGDLMFHMQRQRKLPEEHARFYSAEISLALNYLHERGIIYRDLKLDNVLLDSEGHIKLTDYGMCKEGLRPGDTTSXFCGTPNYIAPEILRGEDYGFSVDWWALGVLMFEMMAGRSPFDIQNTEDYLFQVILEKQIRIPRSLSVKAASVLKSFLNKDPKERLGCHPQTGFADIQGHPFFRNVDWDMMEQKQVVPPFKPNISGEFGLDNFDSQFTNEPVQLXPDDDDIVRKIDQSEFEGFEYINPLRMRPFKRQGSVRR Hydrogen bonds contact Hydrophobic contact | ||||
| 80 | TYK2 tyrosine kinase (TYK2) | 3LXN | 6.44 | |
Target general information Gen name TYK2 Organism Homo sapiens (Human) Uniprot ID TTD ID Synonyms Non-receptor tyrosine-protein kinase TYK2 Protein family Protein kinase superfamily, Tyr protein kinase family, JAK subfamily Biochemical class Kinase Function Probably involved in intracellular signal transduction by being involved in the initiation of type I IFN signaling. Phosphorylates the interferon-alpha/beta receptor alpha chain. Related diseases Immunodeficiency 35 (IMD35) [MIM:611521]: A primary immunodeficiency characterized by recurrent skin abscesses, pneumonia, and highly elevated serum IgE. {ECO:0000269|PubMed:17088085}. The disease is caused by variants affecting the gene represented in this entry. Drugs (DrugBank ID) DB04716; DB14973; DB11817; DB16650; DB12010; DB11763; DB08877; DB08895 Interacts with O95994; Q9ULX6; A8MQ03; Q13643; A6NEM1; P61978; P61978-2; P08238; P17181-1; O60674; Q5VWX1; O75525; Q15323; Q6A162; Q3LI67; Q9NQX1-2; P52630; Q05BL1; Q9BUZ4 EC number EC 2.7.10.2 Uniprot keywords 3D-structure; ATP-binding; Kinase; Nucleotide-binding; Phosphoprotein; Proteomics identification; Reference proteome; Repeat; SH2 domain; Transferase; Tyrosine-protein kinase Protein physicochemical properties Chain ID A Molecular weight (Da) 32807.3 Length 288 Aromaticity 0.11 Instability index 43.07 Isoelectric point 6.37 Charge (pH=7) -3.14 3D Binding mode Sequence DPTVFHKRYLKKIRDLGEGHFGKVSLYCYDPTNDGTGEMVAVKALKADAGPQHRSGWKQEIDILRTLYHEHIIKYKGCCEDAGAASLQLVMEYVPLGSLRDYLPRHSIGLAQLLLFAQQICEGMAYLHAQHYIHRDLAARNVLLDNDRLVKIGDFGLAKAVPEGHEXYRVGDSPVFWYAPECLKEYKFYYASDVWSFGVTLYELLTHCDSSQSPPTKFLELIGIAQGQMTVLRLTELLERGERLPRPDKCPAEVYHLMKNCWETEASFRPTFENLIPILKTVHEKYQG Hydrogen bonds contact Hydrophobic contact | ||||